Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies
- PMID: 36359385
- PMCID: PMC9687899
- DOI: 10.3390/biomedicines10112865
Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies
Abstract
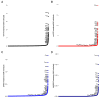
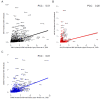
Rare diseases impact up to 400 million individuals globally. Of the thousands of known rare diseases, many are rare neurodevelopmental disorders (RNDDs) impacting children. RNDDs have proven to be difficult to assess epidemiologically for several reasons. The rarity of them makes it difficult to observe them in the population, there is clinical overlap among many disorders, making it difficult to assess the prevalence without genetic testing, and data have yet to be available to have accurate counts of cases. Here, we utilized large sequencing cohorts of individuals with rare, de novo monogenic disorders to estimate the prevalence of variation in over 11,000 genes among cohorts with developmental delay, autism spectrum disorder, and/or epilepsy. We found that the prevalence of many RNDDs is positively correlated to the previously estimated incidence. We identified the most often mutated genes among neurodevelopmental disorders broadly, as well as developmental delay and autism spectrum disorder independently. Finally, we assessed if social media group member numbers may be a valuable way to estimate prevalence. These data are critical for individuals and families impacted by these RNDDs, clinicians and geneticists in their understanding of how common diseases are, and for researchers to potentially prioritize research into particular genes or gene sets.
Keywords: de novo; monogenic; neurodevelopmental disorders; prevalence; rare disease.
Conflict of interest statement
E.E.E. is a scientific advisory board (SAB) member of Variant Bio, Inc. (Seattle, WA, USA).
Figures


References
-
- Rare Disease Act of 2002. [(accessed on 19 May 2022)]; Available online: https://www.congress.gov/107/plaws/publ280/PLAW-107publ280.pdf.
-
- Nguengang Wakap S., Lambert D.M., Olry A., Rodwell C., Gueydan C., Lanneau V., Murphy D., Le Cam Y., Rath A. Estimating Cumulative Point Prevalence of Rare Diseases: Analysis of the Orphanet Database. [(accessed on 24 May 2022)];Eur. J. Hum. Genet. 2020 28:165–173. doi: 10.1038/s41431-019-0508-0. Available online: https://www.nature.com/articles/s41431-019-0508-0. - DOI - PMC - PubMed
-
- Zablotsky B., Black L.I. Prevalence of Children Aged 3–17 Years with Developmental Disabilities, by Urbanicity: United States, 2015–2018. Natl. Health Stat. Report. 2020;139:1–7. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

