Isolation, Characterization, and Genome Analysis of a Novel Bacteriophage, Escherichia Phage vB_EcoM-4HA13, Representing a New Phage Genus in the Novel Phage Family Chaseviridae
- PMID: 36366454
- PMCID: PMC9699118
- DOI: 10.3390/v14112356
Isolation, Characterization, and Genome Analysis of a Novel Bacteriophage, Escherichia Phage vB_EcoM-4HA13, Representing a New Phage Genus in the Novel Phage Family Chaseviridae
Abstract
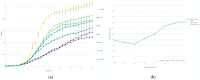
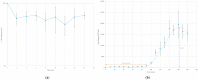
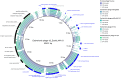
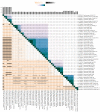
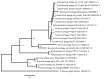
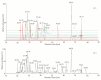
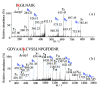
Shiga toxin-producing Escherichia coli (STEC) is one of the leading causes of foodborne illnesses in North America and can lead to severe symptoms, with increased fatality risk for young children. While E. coli O157:H7 remains the dominant STEC serotype associated with foodborne outbreaks, there has been an increasing number of non-O157 STEC outbreaks in recent years. For the food industry, lytic bacteriophages offer an organic, self-limiting alternative to pathogen reduction-one that could replace or reduce the use of chemical and physical food processing methods. From EHEC-enriched sewage, we isolated a novel bacteriophage, vB_EcoM-4HA13 (4HA13). Phenotypic characterizations revealed 4HA13 to possess a myoviral morphotype, with a high specificity to non-motile O111 serotype, and a long latent period (90 min). Through genomic analyses, this 52,401-bp dsDNA phage was found to contain 81 CDS, but no detectable presence of antibiotic resistance, integrase, or virulence genes. A BLASTn search for each of the identified 81 CDS yielded homologues with low levels of similarity. Comparison of RNA polymerase and terminase large subunit amino acid sequences led to the proposal and acceptance of a new bacteriophage family, Chaseviridae, with 4HA13 representing a new species and genus. The discovery of this phage has broadened our current knowledge of bacteriophage diversity.
Keywords: Chaseviridae; Escherichia coli; STEC O111-specific bacteriophage; bacteriophage; genomics; proteomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Characterization of a T4-like Bacteriophage vB_EcoM-Sa45lw as a Potential Biocontrol Agent for Shiga Toxin-Producing Escherichia coli O45 Contaminated on Mung Bean Seeds.Microbiol Spectr. 2022 Feb 23;10(1):e0222021. doi: 10.1128/spectrum.02220-21. Epub 2022 Feb 2. Microbiol Spectr. 2022. PMID: 35107386 Free PMC article.
-
Molecular characterization and safety properties of multi drug-resistant Escherichia coli O157:H7 bacteriophages.BMC Microbiol. 2024 Dec 19;24(1):528. doi: 10.1186/s12866-024-03691-w. BMC Microbiol. 2024. PMID: 39695941 Free PMC article.
-
Genomic, proteomic and physiological characterization of a T5-like bacteriophage for control of Shiga toxin-producing Escherichia coli O157:H7.PLoS One. 2012;7(4):e34585. doi: 10.1371/journal.pone.0034585. Epub 2012 Apr 13. PLoS One. 2012. PMID: 22514640 Free PMC article.
-
Shiga toxin-producing Escherichia coli.Adv Appl Microbiol. 2014;86:145-97. doi: 10.1016/B978-0-12-800262-9.00003-2. Adv Appl Microbiol. 2014. PMID: 24377855 Review.
-
Molecular Modeling the Proteins from the exo-xis Region of Lambda and Shigatoxigenic Bacteriophages.Antibiotics (Basel). 2021 Oct 21;10(11):1282. doi: 10.3390/antibiotics10111282. Antibiotics (Basel). 2021. PMID: 34827220 Free PMC article. Review.
Cited by
-
The Lytic Activity of Bacteriophage ZCSE9 against Salmonella enterica and Its Synergistic Effects with Kanamycin.Viruses. 2023 Mar 31;15(4):912. doi: 10.3390/v15040912. Viruses. 2023. PMID: 37112892 Free PMC article.
-
Newly Isolated Virulent Salmophages for Biocontrol of Multidrug-Resistant Salmonella in Ready-to-Eat Plant-Based Food.Int J Mol Sci. 2023 Jun 14;24(12):10134. doi: 10.3390/ijms241210134. Int J Mol Sci. 2023. PMID: 37373290 Free PMC article.
References
-
- National Outbreak Reporting System (NORS) [(accessed on 12 August 2022)]; Available online: https://wwwn.cdc.gov/norsdashboard/
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

