Expression levels of NONO, a nuclear protein primarily involved in paraspeckles function, are associated with several deregulated molecular pathways and poor clinical outcome in multiple myeloma
- PMID: 36367609
- PMCID: PMC9652193
- DOI: 10.1007/s12672-022-00582-2
Expression levels of NONO, a nuclear protein primarily involved in paraspeckles function, are associated with several deregulated molecular pathways and poor clinical outcome in multiple myeloma
Abstract
Purpose: The NONO protein belongs to the multifunctional family of proteins that can bind DNA, RNA and proteins. It is located in the nucleus of most mammalian cells and can affect almost every step of gene regulation. Dysregulation of NONO has been found in many types of cancer; however, data regarding its expression and relevance in Multiple Myeloma (MM) are virtually absent.
Methods: We took advantage of a large cohort of MM patients enrolled in the Multiple Myeloma Research Foundation CoMMpass study to elucidate better the clinical and biological relevance of NONO expression in the context of the MM genomic landscape and transcriptome.
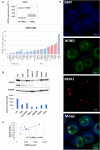
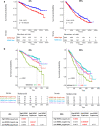
Results: NONO is overexpressed in pathological samples compared to normal controls. In addition, higher NONO expression levels are significant independent prognostic markers of worse clinical outcome in MM. Our results indicate that NONO deregulation may play a pathogenetic role in MM by affecting cell cycle, DNA repair mechanisms, and influencing translation by regulating ribosome biogenesis and assembly. Furthermore, our data suggest NONO involvement in the metabolic reprogramming of glucose metabolism from respiration to aerobic glycolysis, a phenomenon known as the 'Warburg Effect' that supports rapid cancer cell growth, survival, and invasion.
Conclusion: These findings strongly support the need of future investigations for the understanding of the mechanisms of deregulation and the biological role and activity of NONO in MM.
Keywords: Multiple myeloma; NONO; Therapeutic target; Warburg effect.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


References
-
- Saltarella I, Apollonio B, Lamanuzzi A, Desantis V, Mariggio MA, Desaphy JF, Vacca A, Frassanito MA. The landscape of lncRNAs in multiple myeloma: implications in the “Hallmarks of Cancer” clinical perspectives and therapeutic opportunities. Cancers. 2022;14:1963. doi: 10.3390/cancers14081963. - DOI - PMC - PubMed
