Cytoskeletal components can turn wall-less spherical bacteria into kinking helices
- PMID: 36376306
- PMCID: PMC9663586
- DOI: 10.1038/s41467-022-34478-0
Cytoskeletal components can turn wall-less spherical bacteria into kinking helices
Abstract
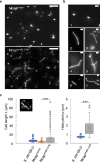
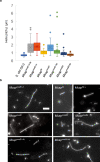
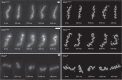
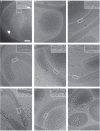
Bacterial cell shape is generally determined through an interplay between the peptidoglycan cell wall and cytoplasmic filaments made of polymerized MreB. Indeed, some bacteria (e.g., Mycoplasma) that lack both a cell wall and mreB genes consist of non-motile cells that are spherical or pleomorphic. However, other members of the same class Mollicutes (e.g., Spiroplasma, also lacking a cell wall) display a helical cell shape and kink-based motility, which is thought to rely on the presence of five MreB isoforms and a specific fibril protein. Here, we show that heterologous expression of Spiroplasma fibril and MreB proteins confers helical shape and kinking ability to Mycoplasma capricolum cells. Isoform MreB5 is sufficient to confer helicity and kink propagation to mycoplasma cells. Cryoelectron microscopy confirms the association of cytoplasmic MreB filaments with the plasma membrane, suggesting a direct effect on membrane curvature. However, in our experiments, the heterologous expression of MreBs and fibril did not result in efficient motility in culture broth, indicating that additional, unknown Spiroplasma components are required for swimming.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
MreB5 Is a Determinant of Rod-to-Helical Transition in the Cell-Wall-less Bacterium Spiroplasma.Curr Biol. 2020 Dec 7;30(23):4753-4762.e7. doi: 10.1016/j.cub.2020.08.093. Epub 2020 Sep 24. Curr Biol. 2020. PMID: 32976813
-
The bacterial linear motor of Spiroplasma melliferum BC3: from single molecules to swimming cells.Mol Microbiol. 2003 Feb;47(3):671-97. doi: 10.1046/j.1365-2958.2003.t01-1-03200.x. Mol Microbiol. 2003. PMID: 12535069
-
Phylogenetic origin and sequence features of MreB from the wall-less swimming bacteria Spiroplasma.Biochem Biophys Res Commun. 2020 Dec 17;533(4):638-644. doi: 10.1016/j.bbrc.2020.09.060. Epub 2020 Oct 13. Biochem Biophys Res Commun. 2020. PMID: 33066960
-
Prospects for the Mechanism of Spiroplasma Swimming.Front Microbiol. 2021 Aug 27;12:706426. doi: 10.3389/fmicb.2021.706426. eCollection 2021. Front Microbiol. 2021. PMID: 34512583 Free PMC article. Review.
-
Exploring Spiroplasma Biology: Opportunities and Challenges.Front Microbiol. 2020 Oct 21;11:589279. doi: 10.3389/fmicb.2020.589279. eCollection 2020. Front Microbiol. 2020. PMID: 33193251 Free PMC article. Review.
Cited by
-
Sequence analyses of a lipoprotein conserved with bacterial actins responsible for swimming motility of wall-less helical Spiroplasma.MicroPubl Biol. 2023 Mar 15;2023:10.17912/micropub.biology.000713. doi: 10.17912/micropub.biology.000713. eCollection 2023. MicroPubl Biol. 2023. PMID: 37033705 Free PMC article.
-
Characterization of heterologously expressed Fibril, a shape and motility determining cytoskeletal protein of the helical bacterium Spiroplasma.iScience. 2022 Sep 3;25(10):105055. doi: 10.1016/j.isci.2022.105055. eCollection 2022 Oct 21. iScience. 2022. PMID: 36157586 Free PMC article.
-
Cloning and sequencing analysis of whole Spiroplasma genome in yeast.Front Microbiol. 2024 May 31;15:1411609. doi: 10.3389/fmicb.2024.1411609. eCollection 2024. Front Microbiol. 2024. PMID: 38881660 Free PMC article.
-
MreB: unraveling the molecular mechanisms of bacterial shape, division, and environmental adaptation.Cell Commun Signal. 2025 Aug 22;23(1):377. doi: 10.1186/s12964-025-02373-y. Cell Commun Signal. 2025. PMID: 40847400 Free PMC article. Review.
-
Integrated Information for Pathogenicity and Treatment of Spiroplasma.Curr Microbiol. 2024 Jul 2;81(8):252. doi: 10.1007/s00284-024-03730-9. Curr Microbiol. 2024. PMID: 38953991 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

