Characterization of Two Novel EF-Hand Proteins Identifies a Clade of Putative Ca2+-Binding Protein Specific to the Ambulacraria
- PMID: 36382242
- PMCID: PMC9648499
- DOI: 10.26502/jbsb.5107030
Characterization of Two Novel EF-Hand Proteins Identifies a Clade of Putative Ca2+-Binding Protein Specific to the Ambulacraria
Abstract
In recent years, transcriptomic databases have become one of the main sources for protein discovery. In our studies of nervous system and digestive tract regeneration in echinoderms, we have identified several transcripts that have attracted our attention. One of these molecules corresponds to a previously unidentified transcript (Orpin) from the sea cucumber Holothuria glaberrima that appeared to be upregulated during intestinal regeneration. We have now identified a second highly similar sequence and analyzed the predicted proteins using bioinformatics tools. Both sequences have EF-hand motifs characteristic of calcium-binding proteins (CaBPs) and N-terminal signal peptides. Sequence comparison analyses such as multiple sequence alignments and phylogenetic analyses only showed significant similarity to sequences from other echinoderms or from hemichordates. Semi-quantitative RT-PCR analyses revealed that transcripts from these sequences are expressed in various tissues including muscle, haemal system, gonads, and mesentery. However, contrary to previous reports, there was no significant differential expression in regenerating tissues. Nonetheless, the identification of unique features in the predicted proteins and their presence in the holothurian draft genome suggest that these might comprise a novel subfamily of EF-hand containing proteins specific to the Ambulacraria clade.
Figures








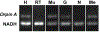


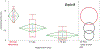

Similar articles
-
Characterization and Expression Analysis of Regeneration-Associated Protein (Aj-Orpin) during Intestinal Regeneration in the Sea Cucumber Apostichopus japonicus.Mar Drugs. 2022 Sep 6;20(9):568. doi: 10.3390/md20090568. Mar Drugs. 2022. PMID: 36135757 Free PMC article.
-
A START-domain-containing protein is a novel marker of nervous system components of the sea cucumber Holothuria glaberrima.Comp Biochem Physiol B Biochem Mol Biol. 2017 Dec;214:57-65. doi: 10.1016/j.cbpb.2017.08.004. Epub 2017 Aug 31. Comp Biochem Physiol B Biochem Mol Biol. 2017. PMID: 28864221 Free PMC article.
-
Distinct profiles of expressed sequence tags during intestinal regeneration in the sea cucumber Holothuria glaberrima.Physiol Genomics. 2007 Oct 22;31(2):203-15. doi: 10.1152/physiolgenomics.00228.2006. Epub 2007 Jun 19. Physiol Genomics. 2007. PMID: 17579180 Free PMC article.
-
Structural and functional diversity of Entamoeba histolytica calcium-binding proteins.Biophys Rev. 2020 Oct 15;12(6):1331-41. doi: 10.1007/s12551-020-00766-6. Online ahead of print. Biophys Rev. 2020. PMID: 33063237 Free PMC article. Review.
-
Calcium binding proteins and calcium signaling in prokaryotes.Cell Calcium. 2015 Mar;57(3):151-65. doi: 10.1016/j.ceca.2014.12.006. Epub 2014 Dec 17. Cell Calcium. 2015. PMID: 25555683 Review.
Cited by
-
Characterization and Expression Analysis of Regeneration-Associated Protein (Aj-Orpin) during Intestinal Regeneration in the Sea Cucumber Apostichopus japonicus.Mar Drugs. 2022 Sep 6;20(9):568. doi: 10.3390/md20090568. Mar Drugs. 2022. PMID: 36135757 Free PMC article.
References
-
- García-Arrarás JE, Estrada-Rodgers L, Santiago R, Torres II, Díaz-Miranda L, Torres-Avillán I. Cellular mechanisms of intestine regeneration in the sea cucumber, Holothuria glaberrima Selenka (Holothuroidea:Echinodermata). J Exp Zool 281 (1998): 288–304. - PubMed
-
- García-Arrarás JE, Greenberg MJ. Visceral Regeneration in Holothurians. Microsc Res Tech. 55(2001): 438–451. - PubMed
-
- Mashanov VS, Zueva OR, García-Arrarás JE. Organization of glial cells in the adult sea cucumber central nervous system. Glia 58(2010): 1581–1593. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
