Brain connectivity meets reservoir computing
- PMID: 36383563
- PMCID: PMC9710781
- DOI: 10.1371/journal.pcbi.1010639
Brain connectivity meets reservoir computing
Abstract
The connectivity of Artificial Neural Networks (ANNs) is different from the one observed in Biological Neural Networks (BNNs). Can the wiring of actual brains help improve ANNs architectures? Can we learn from ANNs about what network features support computation in the brain when solving a task? At a meso/macro-scale level of the connectivity, ANNs' architectures are carefully engineered and such those design decisions have crucial importance in many recent performance improvements. On the other hand, BNNs exhibit complex emergent connectivity patterns at all scales. At the individual level, BNNs connectivity results from brain development and plasticity processes, while at the species level, adaptive reconfigurations during evolution also play a major role shaping connectivity. Ubiquitous features of brain connectivity have been identified in recent years, but their role in the brain's ability to perform concrete computations remains poorly understood. Computational neuroscience studies reveal the influence of specific brain connectivity features only on abstract dynamical properties, although the implications of real brain networks topologies on machine learning or cognitive tasks have been barely explored. Here we present a cross-species study with a hybrid approach integrating real brain connectomes and Bio-Echo State Networks, which we use to solve concrete memory tasks, allowing us to probe the potential computational implications of real brain connectivity patterns on task solving. We find results consistent across species and tasks, showing that biologically inspired networks perform as well as classical echo state networks, provided a minimum level of randomness and diversity of connections is allowed. We also present a framework, bio2art, to map and scale up real connectomes that can be integrated into recurrent ANNs. This approach also allows us to show the crucial importance of the diversity of interareal connectivity patterns, stressing the importance of stochastic processes determining neural networks connectivity in general.
Copyright: © 2022 Damicelli et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




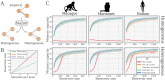

References
-
- Srivastava RK, Greff K, Schmidhuber J. Highway networks. arXiv preprint arXiv:150500387. 2015;.
-
- He K, Zhang X, Ren S, Sun J. Deep Residual Learning for Image Recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2016.
-
- Csordás R, van Steenkiste S, Schmidhuber J. Are Neural Nets Modular? Inspecting Functional Modularity Through Differentiable Weight Masks. 2020.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

