Deep learning to decompose macromolecules into independent Markovian domains
- PMID: 36402768
- PMCID: PMC9675806
- DOI: 10.1038/s41467-022-34603-z
Deep learning to decompose macromolecules into independent Markovian domains
Abstract
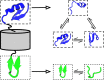
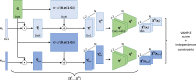
The increasing interest in modeling the dynamics of ever larger proteins has revealed a fundamental problem with models that describe the molecular system as being in a global configuration state. This notion limits our ability to gather sufficient statistics of state probabilities or state-to-state transitions because for large molecular systems the number of metastable states grows exponentially with size. In this manuscript, we approach this challenge by introducing a method that combines our recent progress on independent Markov decomposition (IMD) with VAMPnets, a deep learning approach to Markov modeling. We establish a training objective that quantifies how well a given decomposition of the molecular system into independent subdomains with Markovian dynamics approximates the overall dynamics. By constructing an end-to-end learning framework, the decomposition into such subdomains and their individual Markov state models are simultaneously learned, providing a data-efficient and easily interpretable summary of the complex system dynamics. While learning the dynamical coupling between Markovian subdomains is still an open issue, the present results are a significant step towards learning Ising models of large molecular complexes from simulation data.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Vant, J. W. et al. Protein Structure Prediction 301–315 (Springer, 2020).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

