Chromatin accessibility: methods, mechanisms, and biological insights
- PMID: 36404679
- PMCID: PMC9683059
- DOI: 10.1080/19491034.2022.2143106
Chromatin accessibility: methods, mechanisms, and biological insights
Abstract
Access to DNA is a prerequisite to the execution of essential cellular processes that include transcription, replication, chromosomal segregation, and DNA repair. How the proteins that regulate these processes function in the context of chromatin and its dynamic architectures is an intensive field of study. Over the past decade, genome-wide assays and new imaging approaches have enabled a greater understanding of how access to the genome is regulated by nucleosomes and associated proteins. Additional mechanisms that may control DNA accessibility in vivo include chromatin compaction and phase separation - processes that are beginning to be understood. Here, we review the ongoing development of accessibility measurements, we summarize the different molecular and structural mechanisms that shape the accessibility landscape, and we detail the many important biological functions that are linked to chromatin accessibility.
Keywords: ATAC-seq; Accessibility; HP1; MNase-seq; chromatin; compaction; heterochromatin; linker histones; phase separation; transcription.
Conflict of interest statement
The authors report no potential financial conflict of interest.
Figures
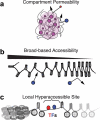

Similar articles
-
HP1 reshapes nucleosome core to promote phase separation of heterochromatin.Nature. 2019 Nov;575(7782):390-394. doi: 10.1038/s41586-019-1669-2. Epub 2019 Oct 16. Nature. 2019. PMID: 31618757 Free PMC article.
-
Determinants and dynamics of genome accessibility.Nat Rev Genet. 2011 Jul 12;12(8):554-64. doi: 10.1038/nrg3017. Nat Rev Genet. 2011. PMID: 21747402 Review.
-
DNA Accessibility by MNase Digestions.Methods Mol Biol. 2018;1689:77-82. doi: 10.1007/978-1-4939-7380-4_7. Methods Mol Biol. 2018. PMID: 29027166
-
Efficient chromatin accessibility mapping in situ by nucleosome-tethered tagmentation.Elife. 2020 Nov 16;9:e63274. doi: 10.7554/eLife.63274. Elife. 2020. PMID: 33191916 Free PMC article.
-
Mechanisms of Nucleosome Dynamics In Vivo.Cold Spring Harb Perspect Med. 2016 Sep 1;6(9):a026666. doi: 10.1101/cshperspect.a026666. Cold Spring Harb Perspect Med. 2016. PMID: 27503998 Free PMC article. Review.
Cited by
-
Transcriptional silencing in Saccharomyces cerevisiae: known unknowns.Epigenetics Chromatin. 2024 Sep 14;17(1):28. doi: 10.1186/s13072-024-00553-7. Epigenetics Chromatin. 2024. PMID: 39272151 Free PMC article. Review.
-
Epigenetic control of multiple genes with a lentiviral vector encoding transcriptional repressors fused to compact zinc finger arrays.Mol Ther Methods Clin Dev. 2024 Apr 24;32(2):101255. doi: 10.1016/j.omtm.2024.101255. eCollection 2024 Jun 13. Mol Ther Methods Clin Dev. 2024. PMID: 38715734 Free PMC article.
-
CXCL Gene Clusters Regulated by Enhancer-Mediated DNA Looping Alteration in Pancreatic Cancer Cells.J Cell Mol Med. 2025 Apr;29(7):e70538. doi: 10.1111/jcmm.70538. J Cell Mol Med. 2025. PMID: 40194986 Free PMC article.
-
Human Endogenous Retroviruses as Novel Therapeutic Targets in Neurodegenerative Disorders.Vaccines (Basel). 2025 Apr 15;13(4):415. doi: 10.3390/vaccines13040415. Vaccines (Basel). 2025. PMID: 40333317 Free PMC article. Review.
-
ScRNA-seq combined with ATAC-seq analysis to explore the metabolic balance mechanism of CCl4-induced liver inflammatory injury.Front Immunol. 2025 Jun 16;16:1600685. doi: 10.3389/fimmu.2025.1600685. eCollection 2025. Front Immunol. 2025. PMID: 40589742 Free PMC article.
References
-
- Klemm SL, Shipony Z, Greenleaf WJ.. Chromatin accessibility and the regulatory epigenome. Nat Rev Genet. 2019;20(4):207–220. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
