Chromatin accessibility: methods, mechanisms, and biological insights
- PMID: 36404679
- PMCID: PMC9683059
- DOI: 10.1080/19491034.2022.2143106
Chromatin accessibility: methods, mechanisms, and biological insights
Abstract
Access to DNA is a prerequisite to the execution of essential cellular processes that include transcription, replication, chromosomal segregation, and DNA repair. How the proteins that regulate these processes function in the context of chromatin and its dynamic architectures is an intensive field of study. Over the past decade, genome-wide assays and new imaging approaches have enabled a greater understanding of how access to the genome is regulated by nucleosomes and associated proteins. Additional mechanisms that may control DNA accessibility in vivo include chromatin compaction and phase separation - processes that are beginning to be understood. Here, we review the ongoing development of accessibility measurements, we summarize the different molecular and structural mechanisms that shape the accessibility landscape, and we detail the many important biological functions that are linked to chromatin accessibility.
Keywords: ATAC-seq; Accessibility; HP1; MNase-seq; chromatin; compaction; heterochromatin; linker histones; phase separation; transcription.
Conflict of interest statement
The authors report no potential financial conflict of interest.
Figures
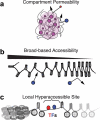

References
-
- Klemm SL, Shipony Z, Greenleaf WJ.. Chromatin accessibility and the regulatory epigenome. Nat Rev Genet. 2019;20(4):207–220. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
