A synthetic genetic array screen for interactions with the RNA helicase DED1 during cell stress in budding yeast
- PMID: 36409020
- PMCID: PMC9836348
- DOI: 10.1093/g3journal/jkac296
A synthetic genetic array screen for interactions with the RNA helicase DED1 during cell stress in budding yeast
Abstract
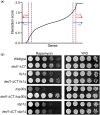
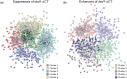
During cellular stress it is essential for cells to alter their gene expression to adapt and survive. Gene expression is regulated at multiple levels, but translation regulation is both a method for rapid changes to the proteome and, as one of the most energy-intensive cellular processes, a way to efficiently redirect cellular resources during stress conditions. Despite this ideal positioning, many of the specifics of how translation is regulated, positively or negatively, during various types of cellular stress remain poorly understood. To further assess this regulation, we examined the essential translation factor Ded1, an RNA helicase that has been previously shown to play important roles in the translational response to cellular stress. In particular, ded1 mutants display an increased resistance to growth inhibition and translation repression induced by the TOR pathway inhibitor, rapamycin, suggesting that normal stress responses are partially defective in these mutants. To gain further insight into Ded1 translational regulation during stress, synthetic genetic array analysis was conducted in the presence of rapamycin with a ded1 mutant and a library of nonessential genes in Saccharomyces cerevisiae to identify positive and negative genetic interactions in an unbiased manner. Here, we report the results of this screen and subsequent network mapping and Gene Ontology-term analysis. Hundreds of candidate interactions were identified, which fell into expected categories, such as ribosomal proteins and amino acid biosynthesis, as well as unexpected ones, including membrane trafficking, sporulation, and protein glycosylation. Therefore, these results provide several specific directions for further comprehensive studies.
Keywords: helicase; rapamycin; stress; translation; yeast.
© The Author(s) 2022. Published by Oxford University Press on behalf of Genetics Society of America.
Conflict of interest statement
None declared.
Figures
Similar articles
-
The RNA Helicase Ded1 Regulates Translation and Granule Formation during Multiple Phases of Cellular Stress Responses.Mol Cell Biol. 2022 Jan 20;42(1):e0024421. doi: 10.1128/MCB.00244-21. Epub 2021 Nov 1. Mol Cell Biol. 2022. PMID: 34723653 Free PMC article.
-
Medulloblastoma-associated mutations in the DEAD-box RNA helicase DDX3X/DED1 cause specific defects in translation.J Biol Chem. 2021 Jan-Jun;296:100296. doi: 10.1016/j.jbc.2021.100296. Epub 2021 Jan 16. J Biol Chem. 2021. PMID: 33460649 Free PMC article.
-
The DEAD-box RNA helicase Ded1 has a role in the translational response to TORC1 inhibition.Mol Biol Cell. 2019 Aug 1;30(17):2171-2184. doi: 10.1091/mbc.E18-11-0702. Epub 2019 May 29. Mol Biol Cell. 2019. PMID: 31141444 Free PMC article.
-
The RNA Helicase Ded1 from Yeast Is Associated with the Signal Recognition Particle and Is Regulated by SRP21.Molecules. 2024 Jun 20;29(12):2944. doi: 10.3390/molecules29122944. Molecules. 2024. PMID: 38931009 Free PMC article.
-
The Ded1/DDX3 subfamily of DEAD-box RNA helicases.Crit Rev Biochem Mol Biol. 2014 Jul-Aug;49(4):343-60. doi: 10.3109/10409238.2014.931339. Crit Rev Biochem Mol Biol. 2014. PMID: 25039764 Review.
Cited by
-
Characterization of RNA Helicase Genes in Ustilago maydis Reveals Links to Stress Response and Teliospore Dormancy.Int J Mol Sci. 2025 Mar 8;26(6):2432. doi: 10.3390/ijms26062432. Int J Mol Sci. 2025. PMID: 40141077 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
