Identification of Novel tet(X3) Variants Resistant To Tigecycline in Acinetobacter Species
- PMID: 36409072
- PMCID: PMC9784759
- DOI: 10.1128/spectrum.01333-22
Identification of Novel tet(X3) Variants Resistant To Tigecycline in Acinetobacter Species
Abstract
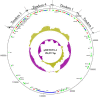
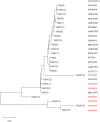
The emergence of the tet(X) gene is a severe challenge to global public health security, as clinical tigecycline resistance shows a rapidly rising trend. In this research, we identified two tigecycline-resistant Acinetobacter sp. strains containing seven novel tet(X3) variants recovered from fecal samples from Chinese farms. The seven Tet(X3) variants showed 15.4% to 99.7% amino acid identity with Tet(X3). By expressing tet(X3.7) and tet(X3.9), the tigecycline MIC values for Escherichia coli JM109 increased 64-fold (from 0.13 to 8 mg/L). However, the other tet(X3) variants did not have a significant change in the MIC of tigecycline. We found that the 26th amino acid site of Tet(X3.7) changed from proline to serine, and the 25th amino acid site of Tet(X3.9) changed from glycine to alanine, which reduced the MIC of tigecycline by 2-fold [the MIC of tet(X3) to tigecycline was 16 mg/L] but did not affect its expression to tigecycline. The tet(X3) variants surrounded by mobile genetic elements appeared in the structure of gene clusters with tandem repeat sequences and were adjacent to the site-specific recombinase-encoding gene xerD. Therefore, there is a risk of horizontal transfer of resistant genes. Our study reports seven novel tet(X3) variants; the continuing emergence of tigecycline variants makes continuous monitoring of resistance to tigecycline even more critical. IMPORTANCE Although it is illegal to use tigecycline and carbapenems to treat bacterial infections in animals, we can still isolate bacteria containing both mobile resistance genes from animals, and tet(X) is currently an essential factor in degrading tigecycline. Here, we characterized two multidrug-resistant Acinetobacter sp. strains that contained vital resistance genes, such as sul2, a blaOXA-164-like gene, floR, tetM, and multiple novel tet(X3) variants with different tandem structures. It is of paramount significance that their mechanism may transfer to other Gram-negative pathogens, even if their tandem structures have no cumulative effect on tigecycline resistance.
Keywords: Acinetobacter schindleri; Acinetobacter variabilis; tet(X); tet(X3); tigecycline resistance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
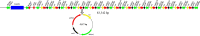



References
-
- Cheng Y, Chen Y, Liu Y, Guo Y, Zhou Y, Xiao T, Zhang S, Xu H, Chen Y, Shan T, Xiao Y, Zhou K. 2020. Identification of novel tetracycline resistance gene tet(X14) and its co-occurrence with tet(X2) in a tigecycline-resistant and colistin-resistant Empedobacter stercoris. Emerg Microbes Infect 9:1843–1852. doi: 10.1080/22221751.2020.1803769. - DOI - PMC - PubMed
-
- Yaghoubi S, Zekiy AO, Krutova M, Gholami M, Kouhsari E, Sholeh M, Ghafouri Z, Maleki F. 2022. Tigecycline antibacterial activity, clinical effectiveness, and mechanisms and epidemiology of resistance: narrative review. Eur J Clin Microbiol Infect Dis 41:1003–1022. doi: 10.1007/s10096-020-04121-1. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

