Mitochondrion-targeted RNA therapies as a potential treatment strategy for mitochondrial diseases
- PMID: 36420220
- PMCID: PMC9664406
- DOI: 10.1016/j.omtn.2022.10.012
Mitochondrion-targeted RNA therapies as a potential treatment strategy for mitochondrial diseases
Abstract
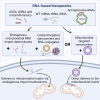
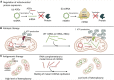
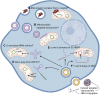
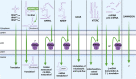
Mitochondrial diseases are one of the largest groups of neurological genetic disorders. Despite continuous efforts of the scientific community, no cure has been developed, and most treatment strategies rely on managing the symptoms. After the success of coronavirus disease 2019 (COVID-19) mRNA vaccines and accelerated US Food and Drug Administration (FDA) approval of four new RNAi drugs, we sought to investigate the potential of mitochondrion-targeting RNA-based therapeutic agents for treatment of mitochondrial diseases. Here we describe the causes and existing therapies for mitochondrial diseases. We then detail potential RNA-based therapeutic strategies for treatment of mitochondrial diseases, including use of antisense oligonucleotides (ASOs) and RNAi drugs, allotopic therapies, and RNA-based antigenomic therapies that aim to decrease the level of deleterious heteroplasmy in affected tissues. Finally, we review different mechanisms by which RNA-based therapeutic agents can be delivered to the mitochondrial matrix, including mitochondrion-targeted nanocarriers and endogenous mitochondrial RNA import pathways.
Keywords: MT: Oligonucleotides, Therapies and Applications; RNA therapeutics; RNA therapy; mitochondrial DNA; mitochondrial RNA import; mitochondrial disease; mitochondrial therapy; mitochondrion-targeted nanocarrier.
© 2022 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Re-Engineering RNA Molecules into Therapeutic Agents.Acc Chem Res. 2019 Apr 16;52(4):1036-1047. doi: 10.1021/acs.accounts.8b00650. Epub 2019 Mar 26. Acc Chem Res. 2019. PMID: 30912917
-
Characterization of chemically modified oligonucleotides targeting a pathogenic mutation in human mitochondrial DNA.Biochimie. 2014 May;100:192-9. doi: 10.1016/j.biochi.2013.08.020. Epub 2013 Aug 28. Biochimie. 2014. PMID: 23994754
-
Transcript-Targeted Therapy Based on RNA Interference and Antisense Oligonucleotides: Current Applications and Novel Molecular Targets.Int J Mol Sci. 2022 Aug 9;23(16):8875. doi: 10.3390/ijms23168875. Int J Mol Sci. 2022. PMID: 36012138 Free PMC article. Review.
-
Therapeutic strategies for mitochondrial disorders.Pediatr Neurol. 2015 Mar;52(3):302-13. doi: 10.1016/j.pediatrneurol.2014.06.023. Epub 2014 Nov 15. Pediatr Neurol. 2015. PMID: 25701186 Review.
-
RNA therapeutics: RNAi and antisense mechanisms and clinical applications.Postdoc J. 2016 Jul;4(7):35-50. doi: 10.14304/surya.jpr.v4n7.5. Postdoc J. 2016. PMID: 27570789 Free PMC article.
Cited by
-
Engineered mitochondria in diseases: mechanisms, strategies, and applications.Signal Transduct Target Ther. 2025 Mar 3;10(1):71. doi: 10.1038/s41392-024-02081-y. Signal Transduct Target Ther. 2025. PMID: 40025039 Free PMC article. Review.
-
Lipid nanoparticle delivery of the CRISPR/Cas9 system directly into the mitochondria of cells carrying m.7778G>T mutation in MtDNA (mt-Atp8).Sci Rep. 2025 Jun 19;15(1):18717. doi: 10.1038/s41598-025-03671-8. Sci Rep. 2025. PMID: 40537532 Free PMC article.
-
Overcoming the Limitations of CRISPR-Cas9 Systems in Saccharomyces cerevisiae: Off-Target Effects, Epigenome, and Mitochondrial Editing.Microorganisms. 2023 Apr 16;11(4):1040. doi: 10.3390/microorganisms11041040. Microorganisms. 2023. PMID: 37110464 Free PMC article. Review.
-
Targeting mitochondria in bone and cartilage diseases: A narrative review.Redox Biol. 2025 Jun;83:103667. doi: 10.1016/j.redox.2025.103667. Epub 2025 May 7. Redox Biol. 2025. PMID: 40354767 Free PMC article. Review.
-
Earlier second polar body transfer and further mitochondrial carryover removal for potential mitochondrial replacement therapy.MedComm (2020). 2023 May 8;4(3):e217. doi: 10.1002/mco2.217. eCollection 2023 Jun. MedComm (2020). 2023. PMID: 37180823 Free PMC article.
References
-
- Rath S., Sharma R., Gupta R., Ast T., Chan C., Durham T.J., Goodman R.P., Grabarek Z., Haas M.E., Hung W.H.W., et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 2021;49:D1541–D1547. doi: 10.1093/nar/gkaa1011. - DOI - PMC - PubMed
-
- Parikh S., Goldstein A., Koenig M.K., Scaglia F., Enns G.M., Saneto R., Anselm I., Cohen B.H., Falk M.J., Greene C., et al. Diagnosis and management of mitochondrial disease: a consensus statement from the Mitochondrial Medicine Society. Genet. Med. 2015;17:689–701. doi: 10.1038/gim.2014.177. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

