A class-mismatched TCR bypasses MHC restriction via an unorthodox but fully functional binding geometry
- PMID: 36424374
- PMCID: PMC9691722
- DOI: 10.1038/s41467-022-34896-0
A class-mismatched TCR bypasses MHC restriction via an unorthodox but fully functional binding geometry
Abstract
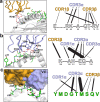
MHC restriction, which describes the binding of TCRs from CD4+ T cells to class II MHC proteins and TCRs from CD8+ T cells to class I MHC proteins, is a hallmark of immunology. Seemingly rare TCRs that break this paradigm exist, but mechanistic insight into their behavior is lacking. TIL1383I is a prototypical class-mismatched TCR, cloned from a CD4+ T cell but recognizing the tyrosinase tumor antigen presented by the class I MHC HLA-A2 in a fully functional manner. Here we find that TIL1383I binds this class I target with a highly atypical geometry. Despite unorthodox binding, TCR signaling, antigen specificity, and the ability to use CD8 are maintained. Structurally, a key feature of TIL1383I is an exceptionally long CDR3β loop that mediates functions that are traditionally performed separately by hypervariable and germline loops in canonical TCR structures. Our findings thus expand the range of known TCR binding geometries compatible with normal function and specificity, provide insight into the determinants of MHC restriction, and may help guide TCR selection and engineering for immunotherapy.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

