Tuning of Gene Expression in Clostridium phytofermentans Using Synthetic Promoters and CRISPRi
- PMID: 36427328
- PMCID: PMC9765743
- DOI: 10.1021/acssynbio.2c00385
Tuning of Gene Expression in Clostridium phytofermentans Using Synthetic Promoters and CRISPRi
Abstract
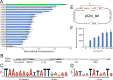
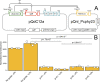
Control of gene expression is fundamental to cell engineering. Here we demonstrate a set of approaches to tune gene expression in Clostridia using the model Clostridium phytofermentans. Initially, we develop a simple benchtop electroporation method that we use to identify a set of replicating plasmids and resistance markers that can be cotransformed into C. phytofermentans. We define a series of promoters spanning a >100-fold expression range by testing a promoter library driving the expression of a luminescent reporter. By insertion of tet operator sites upstream of the reporter, its expression can be quantitatively altered using the Tet repressor and anhydrotetracycline (aTc). We integrate these methods into an aTc-regulated dCas12a system with which we show in vivo CRISPRi-mediated repression of reporter and fermentation genes in C. phytofermentans. Together, these approaches advance genetic transformation and experimental control of gene expression in Clostridia.
Keywords: CRISPRi; Cas12a/Cpf1; Clostridia; electroporation; fermentation; methylome.
Conflict of interest statement
The authors declare no competing financial interest.
Figures




References
-
- Tolonen A. C.; Beauchemin N.; Bayne C.; Li L.; Tan J.; Lee J.; Meehan B. M.; Meisner J.; Millet Y.; LeBlanc G.; Kottler R.; Rapp E.; Murphy C.; Turnbaugh P. J.; von Maltzahn G.; Liu C. M.; van Hylckama Vlieg J. E. T. Synthetic glycans control gut microbiome structure and mitigate colitis in mice. Nat. Commun. 2022, 13, 1244. 10.1038/s41467-022-28856-x. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

