Opposing Roles of IGFBP-3 and Heparanase in Regulating A549 Lung Cancer Cell Survival
- PMID: 36428962
- PMCID: PMC9688904
- DOI: 10.3390/cells11223533
Opposing Roles of IGFBP-3 and Heparanase in Regulating A549 Lung Cancer Cell Survival
Abstract
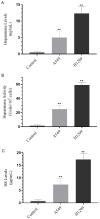
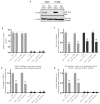
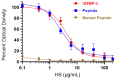
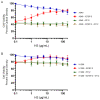
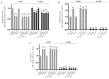
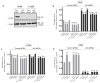
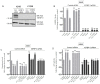
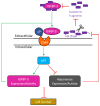
In this study, we examined the roles of heparanase and IGFBP-3 in regulating A549 and H1299 non-small-cell lung cancer (NSCLC) survival. We found that H1299 cells, known to be p53-null with no expression of IGFBP-3, had higher heparanase levels and activity and higher levels of heparan sulfate (HS) in the media compared to the media of A549 cells. Inhibiting heparanase activity or its expression using siRNA had no effect on the levels of IGFBP-3 in the media of A549 cells, reduced the levels of soluble HS fragments, and led to decreased interactions between IGFBP-3 and HS in the media. HS competed with HA for binding to IGFBP-3 or IGFBP-3 peptide (215-KKGFYKKKQCRPSKGRKR-232) but not the mutant peptide (K228AR230A). HS abolished the cytotoxic effects of IGFBP-3 but not upon blocking HA-CD44 signaling with the anti-CD44 antibody (5F12). Blocking HA-CD44 signaling decreased the levels of heparanase in the media of both A549 and H1299 cell lines and increased p53 activity and the levels of IGFBP-3 in A549 cell media. Knockdown of p53 led to increased heparanase levels and reduced IGFBP-3 levels in A549 cell media while knockdown of IGFBP-3 in A549 cells blocked p53 activity and increased heparanase levels in the media.
Keywords: CD44; IGFBP-3; extracellular; heparan sulfate; heparanase; hyaluronan; lung cancer; p53; signaling.
Conflict of interest statement
The authors declare no competing interests.
Figures
References
-
- Siddiqui F., Siddiqui A.H. StatPearls. StatPearls Publishing; Treasure Island, FL, USA: 2021. Lung Cancer.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

