Identification of a Novel GRHL3/HOPX/Wnt/β-Catenin Proto-oncogenic Axis in Squamous Cell Carcinoma of the Esophagus
- PMID: 36442813
- PMCID: PMC10036738
- DOI: 10.1016/j.jcmgh.2022.11.009
Identification of a Novel GRHL3/HOPX/Wnt/β-Catenin Proto-oncogenic Axis in Squamous Cell Carcinoma of the Esophagus
Abstract
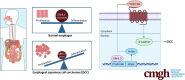
Background & aims: Esophageal squamous cell carcinoma (ESCC) is an aggressive malignancy with a poor long-term prognosis. The molecular mechanisms underlying the initiation and progression of this tumor are largely unknown. The transcription factor GRHL3 functions as a potent tumor suppressor in SCC of skin, head, and neck. This study aims to determine whether GRHL3 also plays a role in the homeostasis of the esophageal epithelium and in the development of ESCC.
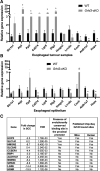
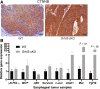
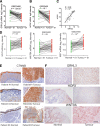
Methods: The effects of Grhl3 deletion on squamous epithelial homeostasis in embryos and adult mice were examined using immunohistochemistry, transmission electron microscopy, and real-time polymerase chain reaction. The conditionally deleted mice were subsequently used to determine susceptibility to ESCC. Whole-transcriptome sequencing (RNA-seq) was performed on ESCC in wild-type and Grhl3 deleted animals. To decipher the signaling pathways, real-time polymerase chain reaction, immunohistochemistry, analysis of chromatin immunoprecipitation sequencing, chromatin immunoprecipitation-polymerase chain reaction, and RNA seq datasets were used. Primary human samples were used to validate the findings in the mouse model.
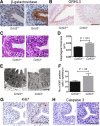
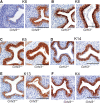
Results: Loss of Grhl3 perturbs the proliferation-differentiation balance in the esophageal epithelium, thereby increasing the susceptibility to esophageal carcinogenesis in adult mice. Grhl3 imparts its tumor suppressor function by regulating the expression of HOPX. We have identified the Wnt/β-catenin pathway as the downstream effectors of GRHL3 and HOPX through our integrated approach using patient-derived ESCC samples and mouse models.
Conclusions: GRHL3 conveys its tumor suppressor function in ESCC through regulating its target gene HOPX, which limits Wnt/β-catenin signaling. Targeted therapies to inhibit this pathway could be a potential treatment strategy for ESCC patients with reduced GRHL3 expression.
Keywords: Cancer Signaling Pathways; RNA Sequencing; Transcriptional Regulation.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Figures




Comment in
-
A Balancing Act: GRHL3 Limits WNT Signaling to Promote Tissue Homeostasis in the Esophageal Epithelium.Cell Mol Gastroenterol Hepatol. 2023;15(5):1247-1248. doi: 10.1016/j.jcmgh.2023.01.011. Epub 2023 Feb 17. Cell Mol Gastroenterol Hepatol. 2023. PMID: 36805430 Free PMC article. No abstract available.
References
-
- Bray F., Ferlay J., Soerjomataram I., et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. - PubMed
-
- Ting S.B., Caddy J., Hislop N., et al. A homolog of Drosophila grainy head is essential for epidermal integrity in mice. Science. 2005;308:411–413. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

