Mechanism of high-mannose N-glycan breakdown and metabolism by Bifidobacterium longum
- PMID: 36443572
- PMCID: PMC10367113
- DOI: 10.1038/s41589-022-01202-4
Mechanism of high-mannose N-glycan breakdown and metabolism by Bifidobacterium longum
Abstract
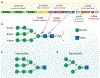
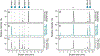
Bifidobacteria are early colonizers of the human gut and play central roles in human health and metabolism. To thrive in this competitive niche, these bacteria evolved the capacity to use complex carbohydrates, including mammalian N-glycans. Herein, we elucidated pivotal biochemical steps involved in high-mannose N-glycan utilization by Bifidobacterium longum. After N-glycan release by an endo-β-N-acetylglucosaminidase, the mannosyl arms are trimmed by the cooperative action of three functionally distinct glycoside hydrolase 38 (GH38) α-mannosidases and a specific GH125 α-1,6-mannosidase. High-resolution cryo-electron microscopy structures revealed that bifidobacterial GH38 α-mannosidases form homotetramers, with the N-terminal jelly roll domain contributing to substrate selectivity. Additionally, an α-glucosidase enables the processing of monoglucosylated N-glycans. Notably, the main degradation product, mannose, is isomerized into fructose before phosphorylation, an unconventional metabolic route connecting it to the bifid shunt pathway. These findings shed light on key molecular mechanisms used by bifidobacteria to use high-mannose N-glycans, a perennial carbon and energy source in the intestinal lumen.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Competing interests
The authors declare no competing interests.
Figures














Similar articles
-
Structure and evolution of the bifidobacterial carbohydrate metabolism proteins and enzymes.Biochem Soc Trans. 2021 Apr 30;49(2):563-578. doi: 10.1042/BST20200163. Biochem Soc Trans. 2021. PMID: 33666221 Free PMC article. Review.
-
Structure and kinetic investigation of Streptococcus pyogenes family GH38 alpha-mannosidase.PLoS One. 2010 Feb 3;5(2):e9006. doi: 10.1371/journal.pone.0009006. PLoS One. 2010. PMID: 20140249 Free PMC article.
-
Recent advances in immobilization strategies for glycosidases.Biotechnol Prog. 2017 Jan;33(1):104-112. doi: 10.1002/btpr.2385. Epub 2016 Oct 31. Biotechnol Prog. 2017. PMID: 27718339 Free PMC article. Review.
-
Fucosyllactose and L-fucose utilization of infant Bifidobacterium longum and Bifidobacterium kashiwanohense.BMC Microbiol. 2016 Oct 26;16(1):248. doi: 10.1186/s12866-016-0867-4. BMC Microbiol. 2016. PMID: 27782805 Free PMC article.
-
Alpha-mannosidases involved in N-glycan processing show cell specificity and distinct subcompartmentalization within the Golgi apparatus of cells in the testis and epididymis.Eur J Cell Biol. 1999 Jul;78(7):441-52. doi: 10.1016/s0171-9335(99)80071-5. Eur J Cell Biol. 1999. PMID: 10472797
Cited by
-
A bayesian approach for parameterizing and predicting plasmid conjugation dynamics.Sci Rep. 2025 Mar 3;15(1):7396. doi: 10.1038/s41598-024-82799-5. Sci Rep. 2025. PMID: 40032848 Free PMC article.
-
Milk glycan metabolism by intestinal bifidobacteria: insights from comparative genomics.Crit Rev Biochem Mol Biol. 2022 Oct-Dec;57(5-6):562-584. doi: 10.1080/10409238.2023.2182272. Epub 2023 Mar 3. Crit Rev Biochem Mol Biol. 2022. PMID: 36866565 Free PMC article. Review.
-
Bifidobacterium longum can degrade the urinary N -glycoprotein Uromodulin.MicroPubl Biol. 2025 Aug 6;2025:10.17912/micropub.biology.001724. doi: 10.17912/micropub.biology.001724. eCollection 2025. MicroPubl Biol. 2025. PMID: 40852033 Free PMC article.
-
Genomic factors shape carbon and nitrogen metabolic niche breadth across Saccharomycotina yeasts.Science. 2024 Apr 26;384(6694):eadj4503. doi: 10.1126/science.adj4503. Epub 2024 Apr 26. Science. 2024. PMID: 38662846 Free PMC article.
-
Priority effects, nutrition and milk glycan-metabolic potential drive Bifidobacterium longum subspecies dynamics in the infant gut microbiome.PeerJ. 2025 Jan 21;13:e18602. doi: 10.7717/peerj.18602. eCollection 2025. PeerJ. 2025. PMID: 39866568 Free PMC article.
References
-
- Andlid TA, D’Aimmo MR & Jastrebova J in The Bifidobacteria and Related Organisms (eds Mattarelli P, Biavati B, Holzapfel WH & Wood BJB) 195–212 (Elsevier, 2018).
-
- Moya-Pérez A, Perez-Villalba A, Benítez-Páez A, Campillo I & Sanz Y Bifidobacterium CECT 7765 modulates early stress-induced immune, neuroendocrine and behavioral alterations in mice. Brain Behav. Immun 65, 43–56 (2017). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

