Temporal phenomic predictions from unoccupied aerial systems can outperform genomic predictions
- PMID: 36445027
- PMCID: PMC9836347
- DOI: 10.1093/g3journal/jkac294
Temporal phenomic predictions from unoccupied aerial systems can outperform genomic predictions
Abstract
A major challenge of genetic improvement and selection is to accurately predict individuals with the highest fitness in a population without direct measurement. Over the last decade, genomic predictions (GP) based on genome-wide markers have become reliable and routine. Now phenotyping technologies, including unoccupied aerial systems (UAS also known as drones), can characterize individuals with a data depth comparable to genomics when used throughout growth. This study, for the first time, demonstrated that the prediction power of temporal UAS phenomic data can achieve or exceed that of genomic data. UAS data containing red-green-blue (RGB) bands over 15 growth time points and multispectral (RGB, red-edge and near infrared) bands over 12 time points were compared across 280 unique maize hybrids. Through cross-validation of untested genotypes in tested environments (CV2), temporal phenomic prediction (TPP), outperformed GP (0.80 vs 0.71); TPP and GP performed similarly in 3 other cross-validation scenarios. Genome-wide association mapping using area under temporal curves of vegetation indices (VIs) revealed 24.5% of a total of 241 discovered loci (59 loci) had associations with multiple VIs, explaining up to 51% of grain yield variation, less than GP and TPP predicted. This suggests TPP, like GP, integrates small effect loci well improving plant fitness predictions. More importantly, TPP appeared to work successfully on unrelated individuals unlike GP.
Keywords: genomic prediction; high-throughput phenotyping; phenomic prediction.
© The Author(s) 2022. Published by Oxford University Press on behalf of Genetics Society of America.
Conflict of interest statement
None declared.
Figures
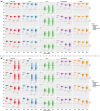


References
-
- Adak A, Murray SC, Anderson SL, Popescu SC, Malambo L, Romay MC, Leon N.. Unoccupied aerial systems discovered overlooked loci capturing the variation of entire growing period in maize. Plant Genome. 2021;14(2):e20102. - PubMed
-
- Adak A, Murray SC, Božinović S, Lindsey R, Nakasagga S, Chatterjee S, Anderson SL, Wilde S.. Temporal vegetation indices and plant height from remotely sensed imagery can predict grain yield and flowering time breeding value in maize via machine learning regression. Remote Sensing. 2021;13(11):2141.
-
- Aguate FM, Trachsel S, González-Pérez L, Burgueño J, Crossa J, Balzarini M, Gouache D, Bogard M, de los Campos G. Use of hyperspectral image data outperforms vegetation indices in prediction of maize yield. Crop Sci. 2017;57:2517–2524.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
