Duck Enteritis Virus Inhibits the cGAS-STING DNA-Sensing Pathway To Evade the Innate Immune Response
- PMID: 36448809
- PMCID: PMC9769366
- DOI: 10.1128/jvi.01578-22
Duck Enteritis Virus Inhibits the cGAS-STING DNA-Sensing Pathway To Evade the Innate Immune Response
Abstract
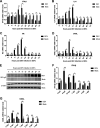
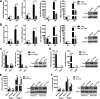
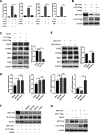
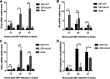
Cyclic GMP-AMP synthase (cGAS), a key DNA sensor, detects cytosolic viral DNA and activates the adaptor protein stimulator of interferon genes (STING) to initiate interferon (IFN) production and host innate antiviral responses. Duck enteritis virus (DEV) is a duck alphaherpesvirus that causes an acute and contagious disease with high mortality in waterfowl. In the present study, we found that DEV inhibits host innate immune responses during the late phase of viral infection. Furthermore, we screened DEV proteins for their ability to inhibit the cGAS-STING DNA-sensing pathway and identified multiple viral proteins, including UL41, US3, UL28, UL53, and UL24, which block IFN-β activation through this pathway. The DEV tegument protein UL41, which exhibited the strongest inhibitory effect, selectively downregulated the expression of interferon regulatory factor 7 (IRF7) by reducing its mRNA accumulation, thereby inhibiting the DNA-sensing pathway. Ectopic expression of UL41 markedly reduced viral DNA-triggered IFN-β production and promoted viral replication, whereas deficiency of UL41 in the context of DEV infection increased the IFN-β response to DEV and suppressed viral replication. In addition, ectopic expression of IRF7 inhibited the replication of the UL41-deficient virus, whereas IRF7 knockdown facilitated its replication. This study is the first report identifying multiple viral proteins encoded by a duck DNA virus, which inhibit the cGAS-STING DNA-sensing pathway. These findings expand our knowledge of DNA sensing in ducks and reveal a mechanism through which DEV antagonizes the host innate immune response. IMPORTANCE Duck enteritis virus (DEV) is a duck alphaherpesvirus that causes an acute and contagious disease with high mortality, resulting in substantial economic losses in the commercial waterfowl industry. The evasion of DNA-sensing pathway-mediated antiviral innate immunity is essential for the persistent infection and replication of many DNA viruses. However, the mechanisms used by DEV to modulate the DNA-sensing pathway remain poorly understood. In the present study, we found that DEV encodes multiple viral proteins to inhibit the cGAS-STING DNA-sensing pathway. The DEV tegument protein UL41 selectively diminished the accumulation of interferon regulatory factor 7 (IRF7) mRNA, thereby inhibiting the DNA-sensing pathway. Loss of UL41 potently enhanced the IFN-β response to DEV and impaired viral replication in ducks. These findings provide insights into the host-virus interaction during DEV infection and help develop new live attenuated vaccines against DEV.
Keywords: DNA sensing; cGAS-STING; duck enteritis virus; innate immunity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Duck Enteritis Virus Protein Kinase US3 Inhibits DNA Sensing Signaling by Phosphorylating Interferon Regulatory Factor 7.Microbiol Spectr. 2022 Dec 21;10(6):e0229922. doi: 10.1128/spectrum.02299-22. Epub 2022 Oct 26. Microbiol Spectr. 2022. PMID: 36287016 Free PMC article.
-
Herpes Simplex Virus 1 Abrogates the cGAS/STING-Mediated Cytosolic DNA-Sensing Pathway via Its Virion Host Shutoff Protein, UL41.J Virol. 2017 Feb 28;91(6):e02414-16. doi: 10.1128/JVI.02414-16. Print 2017 Mar 15. J Virol. 2017. PMID: 28077645 Free PMC article.
-
Inhibition of DNA-Sensing Pathway by Marek's Disease Virus VP23 Protein through Suppression of Interferon Regulatory Factor 7 Activation.J Virol. 2019 Feb 5;93(4):e01934-18. doi: 10.1128/JVI.01934-18. Print 2019 Feb 15. J Virol. 2019. PMID: 30518647 Free PMC article.
-
How Does African Swine Fever Virus Evade the cGAS-STING Pathway?Pathogens. 2024 Nov 2;13(11):957. doi: 10.3390/pathogens13110957. Pathogens. 2024. PMID: 39599510 Free PMC article. Review.
-
How Dengue Virus Circumvents Innate Immunity.Front Immunol. 2018 Dec 4;9:2860. doi: 10.3389/fimmu.2018.02860. eCollection 2018. Front Immunol. 2018. PMID: 30564245 Free PMC article. Review.
Cited by
-
The role of the cGAS-STING pathway in metabolic diseases.Heliyon. 2024 Jun 14;10(12):e33093. doi: 10.1016/j.heliyon.2024.e33093. eCollection 2024 Jun 30. Heliyon. 2024. PMID: 38988528 Free PMC article. Review.
-
An alpha-herpesvirus employs host HEXIM1 to promote viral transcription.J Virol. 2024 Mar 19;98(3):e0139223. doi: 10.1128/jvi.01392-23. Epub 2024 Feb 16. J Virol. 2024. PMID: 38363111 Free PMC article.
-
Development and immunogenicity evaluation of a quadruple-gene-deleted pseudorabies virus strain.Front Microbiol. 2024 Sep 20;15:1479794. doi: 10.3389/fmicb.2024.1479794. eCollection 2024. Front Microbiol. 2024. PMID: 39372271 Free PMC article.
-
IRF7: role and regulation in immunity and autoimmunity.Front Immunol. 2023 Aug 10;14:1236923. doi: 10.3389/fimmu.2023.1236923. eCollection 2023. Front Immunol. 2023. PMID: 37638030 Free PMC article. Review.
-
Duck plague virus UL24 protein initiates K48/K63-linked IRF7 polyubiquitination to antagonize the innate immune response.Poult Sci. 2024 Dec;103(12):104378. doi: 10.1016/j.psj.2024.104378. Epub 2024 Oct 4. Poult Sci. 2024. PMID: 39418790 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

