Cheminformatics analysis of chemicals that increase estrogen and progesterone synthesis for a breast cancer hazard assessment
- PMID: 36450809
- PMCID: PMC9712655
- DOI: 10.1038/s41598-022-24889-w
Cheminformatics analysis of chemicals that increase estrogen and progesterone synthesis for a breast cancer hazard assessment
Abstract
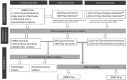
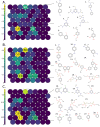
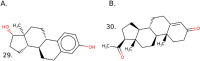
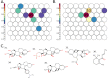
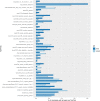
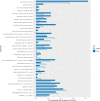
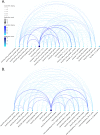
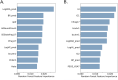
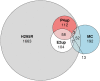
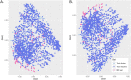
Factors that increase estrogen or progesterone (P4) action are well-established as increasing breast cancer risk, and many first-line treatments to prevent breast cancer recurrence work by blocking estrogen synthesis or action. In previous work, using data from an in vitro steroidogenesis assay developed for the U.S. Environmental Protection Agency (EPA) ToxCast program, we identified 182 chemicals that increased estradiol (E2up) and 185 that increased progesterone (P4up) in human H295R adrenocortical carcinoma cells, an OECD validated assay for steroidogenesis. Chemicals known to induce mammary effects in vivo were very likely to increase E2 or P4 synthesis, further supporting the importance of these pathways for breast cancer. To identify additional chemical exposures that may increase breast cancer risk through E2 or P4 steroidogenesis, we developed a cheminformatics approach to identify structural features associated with these activities and to predict other E2 or P4 steroidogens from their chemical structures. First, we used molecular descriptors and physicochemical properties to cluster the 2,012 chemicals screened in the steroidogenesis assay using a self-organizing map (SOM). Structural features such as triazine, phenol, or more broadly benzene ramified with halide, amine or alcohol, are enriched for E2 or P4up chemicals. Among E2up chemicals, phenol and benzenone are found as significant substructures, along with nitrogen-containing biphenyls. For P4up chemicals, phenol and complex aromatic systems ramified with oxygen-based groups such as flavone or phenolphthalein are significant substructures. Chemicals that are active for both E2up and P4up are enriched with substructures such as dihydroxy phosphanedithione or are small chemicals that contain one benzene ramified with chlorine, alcohol, methyl or primary amine. These results are confirmed with a chemotype ToxPrint analysis. Then, we used machine learning and artificial intelligence algorithms to develop and validate predictive classification QSAR models for E2up and P4up chemicals. These models gave reasonable external prediction performances (balanced accuracy ~ 0.8 and Matthews Coefficient Correlation ~ 0.5) on an external validation. The QSAR models were enriched by adding a confidence score that considers the chemical applicability domain and a ToxPrint assessment of the chemical. This profiling and these models may be useful to direct future testing and risk assessments for chemicals related to breast cancer and other hormonally-mediated outcomes.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

