DDX17 helicase promotes resolution of R-loop-mediated transcription-replication conflicts in human cells
- PMID: 36453994
- PMCID: PMC9757067
- DOI: 10.1093/nar/gkac1116
DDX17 helicase promotes resolution of R-loop-mediated transcription-replication conflicts in human cells
Abstract
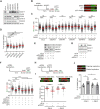
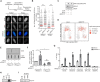
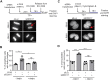
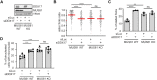
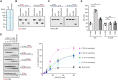
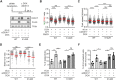
R-loops are three-stranded nucleic acid structures composed of an RNA:DNA hybrid and displaced DNA strand. These structures can halt DNA replication when formed co-transcriptionally in the opposite orientation to replication fork progression. A recent study has shown that replication forks stalled by co-transcriptional R-loops can be restarted by a mechanism involving fork cleavage by MUS81 endonuclease, followed by ELL-dependent reactivation of transcription, and fork religation by the DNA ligase IV (LIG4)/XRCC4 complex. However, how R-loops are eliminated to allow the sequential restart of transcription and replication in this pathway remains elusive. Here, we identified the human DDX17 helicase as a factor that associates with R-loops and counteracts R-loop-mediated replication stress to preserve genome stability. We show that DDX17 unwinds R-loops in vitro and promotes MUS81-dependent restart of R-loop-stalled forks in human cells in a manner dependent on its helicase activity. Loss of DDX17 helicase induces accumulation of R-loops and the formation of R-loop-dependent anaphase bridges and micronuclei. These findings establish DDX17 as a component of the MUS81-LIG4-ELL pathway for resolution of R-loop-mediated transcription-replication conflicts, which may be involved in R-loop unwinding.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


References
-
- Gaillard H., Garcia-Muse T., Aguilera A.. Replication stress and cancer. Nat. Rev. Cancer. 2015; 15:276–289. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

