Pleiotropic fitness effects of a Drosophila odorant-binding protein
- PMID: 36454098
- PMCID: PMC9911060
- DOI: 10.1093/g3journal/jkac307
Pleiotropic fitness effects of a Drosophila odorant-binding protein
Abstract
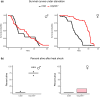
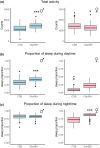
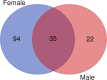
Insect odorant-binding proteins (OBPs) are members of a rapidly evolving multigene family traditionally thought to facilitate chemosensation. However, studies on Drosophila have shown that members of this family have evolved functions beyond chemosensation, as evident from their expression in reproductive tissues and the brain. Previous studies implicated diverse functions of Obp56h, a member of the largest gene cluster of the D. melanogaster Obp repertoire. Here, we examined the effect of CRISPR/Cas9-mediated deletion of Obp56h on 2 fitness phenotypes, on resistance to starvation stress and heat stress, and on locomotion and sleep phenotypes. Obp56h-/- mutants show a strong sexually dimorphic effect on starvation stress survival, with females being more resistant to starvation stress than the control. In contrast, Obp56h-/- females, but not males, are highly sensitive to heat stress. Both sexes show changes in locomotion and sleep patterns. Transcriptional profiling of RNA from heads of Obp56h-/- flies and the wildtype control reveals differentially expressed genes, including gene products associated with antimicrobial immune responses and members of the Turandot family of stress-induced secreted peptides. In addition, differentially expressed genes of unknown function were identified in both sexes. Genes encoding components of the mitochondrial electron transport chain, cuticular proteins, gene products associated with regulation of feeding behavior (Lst and CCHa2), ribosomal proteins, lncRNAs, snoRNAs, tRNAs, and snRNAs show changes in transcript abundances in Obp56h-/- females. These differentially expressed genes are likely to contribute to Obp56h-mediated effects on the diverse phenotypes that arise upon deletion of this OBP.
Keywords: Obp56h; CRISPR-Cas9; RNA sequencing; chemosensation; fitness; heat stress; pleiotropy; sexual dimorphism; sleep; starvation resistance.
© The Author(s) 2022. Published by Oxford University Press on behalf of the Genetics Society of America.
Conflict of interest statement
Conflicts of interest The authors declare no conflict of interest.
Figures

Similar articles
-
Obp56h Modulates Mating Behavior in Drosophila melanogaster.G3 (Bethesda). 2016 Oct 13;6(10):3335-3342. doi: 10.1534/g3.116.034595. G3 (Bethesda). 2016. PMID: 27558663 Free PMC article.
-
Functional Diversification, Redundancy, and Epistasis among Paralogs of the Drosophila melanogaster Obp50a-d Gene Cluster.Mol Biol Evol. 2021 May 4;38(5):2030-2044. doi: 10.1093/molbev/msab004. Mol Biol Evol. 2021. PMID: 33560417 Free PMC article.
-
Trans regulation of an odorant binding protein by a proto-Y chromosome affects male courtship in house fly.Elife. 2024 Oct 18;13:e90349. doi: 10.7554/eLife.90349. Elife. 2024. PMID: 39422654 Free PMC article.
-
Evolution of expression patterns of two odorant-binding protein genes, Obp57d and Obp57e, in Drosophila.Gene. 2010 Nov 1;467(1-2):25-34. doi: 10.1016/j.gene.2010.07.006. Epub 2010 Jul 15. Gene. 2010. PMID: 20637846
-
The 40-Year Mystery of Insect Odorant-Binding Proteins.Biomolecules. 2021 Mar 30;11(4):509. doi: 10.3390/biom11040509. Biomolecules. 2021. PMID: 33808208 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
