Mutation of gdpS gene induces a viable but non-culturable state in Staphylococcus epidermidis and changes in the global transcriptional profile
- PMID: 36457079
- PMCID: PMC9714401
- DOI: 10.1186/s12866-022-02708-6
Mutation of gdpS gene induces a viable but non-culturable state in Staphylococcus epidermidis and changes in the global transcriptional profile
Abstract
Background: In the genome of staphylococci, only the gdpS gene encodes the conserved GGDEF domain, which is the characteristic of diguanylate cyclases. In our previous study, we have demonstrated that the gdpS gene can modulate biofilm formation by positively regulating the expression of ica operon in Staphylococcus epidermidis. Moreover, this regulation seems to be independent of the c-di-GMP signaling pathway and the protein-coding function of this gene. Therefore, the biological function of the gdpS gene remains to be further investigated.
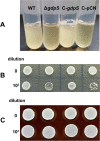
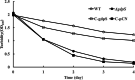
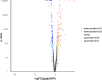
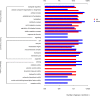
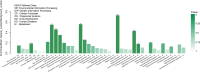
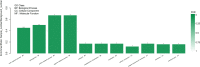
Results: In the present study, it was observed that mutation of the gdpS gene induced S. epidermidis to enter into a presumed viable but nonculturable state (VBNC) after cryopreservation with glycerol. Similarly, when moved from liquid to solid culture medium, the gdpS mutant strain also exhibited a VBNC state. Compared with the wild-type strain, the gdpS mutant strain autolyzed more quickly during storage at 4℃, indicating its increased susceptibility to low temperature. Transcriptional profiling analysis showed that the gdpS mutation affected the transcription of 188 genes (92 genes were upregulated and 96 genes were downregulated). Specifically, genes responsible for glycerol metabolism were most markedly upregulated and most of the altered genes in the mutant strain are those involved in nitrogen metabolism. In addition, the most significantly downregulated genes included the betB gene, whose product catalyzes the synthesis of glycine betaine and confers tolerance to cold.
Conclusion: The preliminary results suggest that the gdpS gene may participate in VBNC formation of S. epidermidis in face of adverse environmental factors, which is probably achieved by regulating expression of energy metabolism genes. Besides, the gdpS gene is critical for S. epidermidis to survive low temperature, and the underlying mechanism may be partly explained by its influence on expression of betB gene.
Keywords: Low temperature; Staphylococcus epidermidis; Transcriptional profile; Viable but nonculturable; gdpS.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

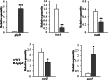
Similar articles
-
A staphylococcal GGDEF domain protein regulates biofilm formation independently of cyclic dimeric GMP.J Bacteriol. 2008 Aug;190(15):5178-89. doi: 10.1128/JB.00375-08. Epub 2008 May 23. J Bacteriol. 2008. PMID: 18502872 Free PMC article.
-
The Staphylococcus epidermidis gdpS regulates biofilm formation independently of its protein-coding function.Microb Pathog. 2017 Apr;105:264-271. doi: 10.1016/j.micpath.2017.02.045. Epub 2017 Mar 1. Microb Pathog. 2017. PMID: 28259672
-
The Vancomycin Resistance-Associated Regulatory System VraSR Modulates Biofilm Formation of Staphylococcus epidermidis in an ica-Dependent Manner.mSphere. 2021 Oct 27;6(5):e0064121. doi: 10.1128/mSphere.00641-21. Epub 2021 Sep 22. mSphere. 2021. PMID: 34550006 Free PMC article.
-
codY and pdhA Expression Is Induced in Staphylococcus epidermidis Biofilm and Planktonic Populations With Higher Proportions of Viable but Non-Culturable Cells.Front Cell Infect Microbiol. 2021 Nov 15;11:771666. doi: 10.3389/fcimb.2021.771666. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 34869073 Free PMC article.
-
Genetic regulation of the intercellular adhesion locus in staphylococci.Front Cell Infect Microbiol. 2012 Mar 26;2:38. doi: 10.3389/fcimb.2012.00038. eCollection 2012. Front Cell Infect Microbiol. 2012. PMID: 23061050 Free PMC article. Review.
Cited by
-
SarZ inhibits the hemolytic activity through regulation of phenol soluble modulins in Staphylococcus epidermidis.Front Cell Infect Microbiol. 2024 Nov 19;14:1476287. doi: 10.3389/fcimb.2024.1476287. eCollection 2024. Front Cell Infect Microbiol. 2024. PMID: 39628668 Free PMC article.
-
Mechanisms underlying the low-temperature adaptation of 17β-estradiol-degrading bacterial strain Rhodococcus sp. RCBS9: insights from physiological and transcriptomic analyses.Front Microbiol. 2024 Nov 21;15:1465627. doi: 10.3389/fmicb.2024.1465627. eCollection 2024. Front Microbiol. 2024. PMID: 39640852 Free PMC article.
-
Cronobacter sakazakii lysozyme inhibitor LprI mediated by HmsP and c-di-GMP is essential for biofilm formation and virulence.Appl Environ Microbiol. 2024 Oct 23;90(10):e0156424. doi: 10.1128/aem.01564-24. Epub 2024 Sep 19. Appl Environ Microbiol. 2024. PMID: 39297664 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- KJ2020A0602/the Key Program of Educational Commission of Anhui Province
- FDMV-2020005/Open Research Fund Program of Key Laboratory of Medical Molecular Virology (MOE/NHC), and Fudan University
- 81871622/National Natural Science Foundation of China
- 82072249/National Natural Science Foundation of China
- 81991532/National Natural Science Foundation of China
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

