Shared graft-versus-leukemia minor histocompatibility antigens in DISCOVeRY-BMT
- PMID: 36477467
- PMCID: PMC10182302
- DOI: 10.1182/bloodadvances.2022008863
Shared graft-versus-leukemia minor histocompatibility antigens in DISCOVeRY-BMT
Abstract
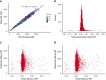
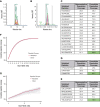
T-cell responses to minor histocompatibility antigens (mHAs) mediate graft-versus-leukemia (GVL) effects and graft-versus-host disease (GVHD) in allogeneic hematopoietic cell transplantation. Therapies that boost T-cell responses improve allogeneic hematopoietic cell transplant (alloHCT) efficacy but are limited by concurrent increases in the incidence and severity of GVHD. mHAs with expression restricted to hematopoietic tissue (GVL mHAs) are attractive targets for driving GVL without causing GVHD. Prior work to identify mHAs has focused on a small set of mHAs or population-level single-nucleotide polymorphism-association studies. We report the discovery of a large set of novel GVL mHAs based on predicted immunogenicity, tissue expression, and degree of sharing among donor-recipient pairs (DRPs) in the DISCOVeRY-BMT data set of 3231 alloHCT DRPs. The total number of predicted mHAs varied by HLA allele, and the total number and number of each class of mHA significantly differed by recipient genomic ancestry group. From the pool of predicted mHAs, we identified the smallest sets of GVL mHAs needed to cover 100% of DRPs with a given HLA allele. We used mass spectrometry to search for high-population frequency mHAs for 3 common HLA alleles. We validated 24 predicted novel GVL mHAs that are found cumulatively within 98.8%, 60.7%, and 78.9% of DRPs within DISCOVeRY-BMT that express HLA-A∗02:01, HLA-B∗35:01, and HLA-C∗07:02, respectively. We confirmed the immunogenicity of an example novel mHA via T-cell coculture with peptide-pulsed dendritic cells. This work demonstrates that the identification of shared mHAs is a feasible and promising technique for expanding mHA-targeting immunotherapeutics.
© 2023 by The American Society of Hematology. Licensed under Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International (CC BY-NC-ND 4.0), permitting only noncommercial, nonderivative use with attribution. All other rights reserved.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures






References
-
- Mullally A, Ritz J. Beyond HLA: the significance of genomic variation for allogeneic hematopoietic stem cell transplantation. Blood. 2006;109(4):1355–1362. - PubMed
-
- Horowitz MM, Gale RP, Sondel PM, et al. Graft-versus-leukemia reactions after bone marrow transplantation. Blood. 1990;75(3):555–562. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

