Displacement and dissociation of oligonucleotides during DNA hairpin closure under strain
- PMID: 36478056
- PMCID: PMC9757040
- DOI: 10.1093/nar/gkac1113
Displacement and dissociation of oligonucleotides during DNA hairpin closure under strain
Abstract
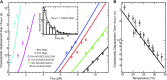
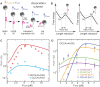
The hybridization kinetic of an oligonucleotide to its template is a fundamental step in many biological processes such as replication arrest, CRISPR recognition, DNA sequencing, DNA origami, etc. Although single kinetic descriptions exist for special cases of this problem, there are no simple general prediction schemes. In this work, we have measured experimentally, with no fluorescent labelling, the displacement of an oligonucleotide from its substrate in two situations: one corresponding to oligonucleotide binding/unbinding on ssDNA and one in which the oligonucleotide is displaced by the refolding of a dsDNA fork. In this second situation, the fork is expelling the oligonucleotide thus significantly reducing its residence time. To account for our data in these two situations, we have constructed a mathematical model, based on the known nearest neighbour dinucleotide free energies, and provided a good estimate of the residence times of different oligonucleotides (DNA, RNA, LNA) of various lengths in different experimental conditions (force, temperature, buffer conditions, presence of mismatches, etc.). This study provides a foundation for the dynamics of oligonucleotide displacement, a process of importance in numerous biological and bioengineering contexts.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




References
-
- Shin J.-S., Pierce N.A.. A synthetic DNA walker for molecular transport. J. Am. Chem. Soc. 2004; 126:10834–10835. - PubMed
-
- Winfree E., Liu F., Wenzler L.A., Seeman N.C.. Design and self-assembly of two-dimensional DNA crystals. Nature. 1998; 394:539–544. - PubMed
-
- Zhang D.Y., Seelig G.. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 2011; 3:103–113. - PubMed

