Bayesian model and selection signature analyses reveal risk factors for canine atopic dermatitis
- PMID: 36482174
- PMCID: PMC9731970
- DOI: 10.1038/s42003-022-04279-8
Bayesian model and selection signature analyses reveal risk factors for canine atopic dermatitis
Abstract
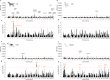
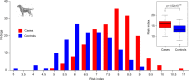
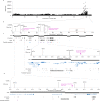
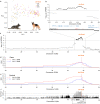
Canine atopic dermatitis is an inflammatory skin disease with clinical similarities to human atopic dermatitis. Several dog breeds are at increased risk for developing this disease but previous genetic associations are poorly defined. To identify additional genetic risk factors for canine atopic dermatitis, we here apply a Bayesian mixture model adapted for mapping complex traits and a cross-population extended haplotype test to search for disease-associated loci and selective sweeps in four dog breeds at risk for atopic dermatitis. We define 15 associated loci and eight candidate regions under selection by comparing cases with controls. One associated locus is syntenic to the major genetic risk locus (Filaggrin locus) in human atopic dermatitis. One selection signal in common type Labrador retriever cases positions across the TBC1D1 gene (body weight) and one signal of selection in working type German shepherd controls overlaps the LRP1B gene (brain), near the KYNU gene (psoriasis). In conclusion, we identify candidate genes, including genes belonging to the same biological pathways across multiple loci, with potential relevance to the pathogenesis of canine atopic dermatitis. The results show genetic similarities between dog and human atopic dermatitis, and future across-species genetic comparisons are hereby further motivated.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

