Robust induction of primordial germ cells of white rhinoceros on the brink of extinction
- PMID: 36490332
- PMCID: PMC9733929
- DOI: 10.1126/sciadv.abp9683
Robust induction of primordial germ cells of white rhinoceros on the brink of extinction
Abstract
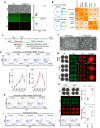
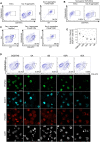
In vitro gametogenesis, the process of generating gametes from pluripotent cells in culture, is a powerful tool for improving our understanding of germ cell development and an alternative source of gametes. Here, we induced primordial germ cell-like cells (PGCLCs) from pluripotent stem cells of the northern white rhinoceros (NWR), a species for which only two females remain, and southern white rhinoceros (SWR), the closest species to the NWR. PGCLC differentiation from SWR embryonic stem cells is highly reliant on bone morphogenetic protein and WNT signals. Genetic analysis revealed that SRY-box transcription factor 17 (SOX17) is essential for SWR-PGCLC induction. Under the defined condition, NWR induced pluripotent stem cells differentiated into PGCLCs. We also identified cell surface markers, CD9 and Integrin subunit alpha 6 (ITGA6), that enabled us to isolate PGCLCs without genetic alteration in pluripotent stem cells. This study provides a first step toward the production of NWR gametes in culture and understanding of the basic mechanism of primordial germ cell specification in a large animal.
Figures





References
-
- A. D. Barnosky, N. Matzke, S. Tomiya, G. O. U. Wogan, B. Swartz, T. B. Quental, C. Marshall, J. L. McGuire, E. L. Lindsey, K. C. Maguire, B. Mersey, E. A. Ferrer,Has the Earth’s sixth mass extinction already arrived? Nature 471,51–57 (2011). - PubMed
-
- R. H. Emslie, M. Brooks, in African Rhino: Status Survey and Conservation Action Plan (IUCN, 1999), pp. 1–92.
-
- K. Hillman Smith, J. A. Ndey,Post-war effects on the rhinos and elephants of Garamba National Park. Pachyderm 39,106–110 (2005).
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

