The draft genome of Andean Rhodopseudomonas sp. strain AZUL predicts genome plasticity and adaptation to chemical homeostasis
- PMID: 36494611
- PMCID: PMC9733117
- DOI: 10.1186/s12866-022-02685-w
The draft genome of Andean Rhodopseudomonas sp. strain AZUL predicts genome plasticity and adaptation to chemical homeostasis
Abstract
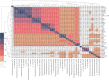
The genus Rhodopseudomonas comprises purple non-sulfur bacteria with extremely versatile metabolisms. Characterization of several strains revealed that each is a distinct ecotype highly adapted to its specific micro-habitat. Here we present the sequencing, genomic comparison and functional annotation of AZUL, a Rhodopseudomonas strain isolated from a high altitude Andean lagoon dominated by extreme conditions and fluctuating levels of chemicals. Average nucleotide identity (ANI) analysis of 39 strains of this genus showed that the genome of AZUL is 96.2% identical to that of strain AAP120, which suggests that they belong to the same species. ANI values also show clear separation at the species level with the rest of the strains, being more closely related to R. palustris. Pangenomic analyses revealed that the genus Rhodopseudomonas has an open pangenome and that its core genome represents roughly 5 to 12% of the total gene repertoire of the genus. Functional annotation showed that AZUL has genes that participate in conferring genome plasticity and that, in addition to sharing the basal metabolic complexity of the genus, it is also specialized in metal and multidrug resistance and in responding to nutrient limitation. Our results also indicate that AZUL might have evolved to use some of the mechanisms involved in resistance as redox reactions for bioenergetic purposes. Most of those features are shared with strain AAP120, and mainly involve the presence of additional orthologs responsible for the mentioned processes. Altogether, our results suggest that AZUL, one of the few bacteria from its habitat with a sequenced genome, is highly adapted to the extreme and changing conditions that constitute its niche.
Keywords: Chemical resistance; High-altitude Andean lakes; Pangenomic analysis; Purple non-sulfur bacteria; Rhodopseudomonas.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




Similar articles
-
Genomic and genetic sequence information of strains assigned to the genus Rhodopseudomonas reveal the great heterogeneity of the group and identify strain Rhodopseudomonas palustris DSM 123T as the authentic type strain of this species.Int J Syst Evol Microbiol. 2020 Jun;70(6):3932-3938. doi: 10.1099/ijsem.0.004077. Epub 2020 Jun 3. Int J Syst Evol Microbiol. 2020. PMID: 32496176
-
Genomic and Phylogenetic Characterization of Rhodopseudomonas infernalis sp. nov., Isolated from the Hell Creek Watershed (Nebraska).Microorganisms. 2022 Oct 13;10(10):2024. doi: 10.3390/microorganisms10102024. Microorganisms. 2022. PMID: 36296300 Free PMC article.
-
Multiple genome sequences reveal adaptations of a phototrophic bacterium to sediment microenvironments.Proc Natl Acad Sci U S A. 2008 Nov 25;105(47):18543-8. doi: 10.1073/pnas.0809160105. Epub 2008 Nov 19. Proc Natl Acad Sci U S A. 2008. PMID: 19020098 Free PMC article.
-
Descriptions of Rhodopseudomonas parapalustris sp. nov., Rhodopseudomonas harwoodiae sp. nov. and Rhodopseudomonas pseudopalustris sp. nov., and emended description of Rhodopseudomonas palustris.Int J Syst Evol Microbiol. 2012 Aug;62(Pt 8):1790-1798. doi: 10.1099/ijs.0.026815-0. Epub 2011 Oct 10. Int J Syst Evol Microbiol. 2012. PMID: 21986724
-
Rhodopseudomonas palustris: A biotechnology chassis.Biotechnol Adv. 2022 Nov;60:108001. doi: 10.1016/j.biotechadv.2022.108001. Epub 2022 Jun 6. Biotechnol Adv. 2022. PMID: 35680002 Review.
Cited by
-
A Rhodopseudomonas strain with a substantially smaller genome retains the core metabolic versatility of its genus.Appl Environ Microbiol. 2025 Apr 23;91(4):e0205624. doi: 10.1128/aem.02056-24. Epub 2025 Mar 10. Appl Environ Microbiol. 2025. PMID: 40062894 Free PMC article.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

