Methodological challenges in the genomic analysis of an endangered mammal population with low genetic diversity
- PMID: 36496459
- PMCID: PMC9741620
- DOI: 10.1038/s41598-022-25619-y
Methodological challenges in the genomic analysis of an endangered mammal population with low genetic diversity
Abstract
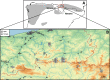
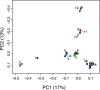
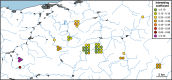
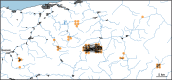
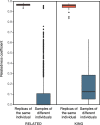
Recently, populations of various species with very low genetic diversity have been discovered. Some of these persist in the long term, but others could face extinction due to accelerated loss of fitness. In this work, we characterize 45 individuals of one of these populations, belonging to the Iberian desman (Galemys pyrenaicus). For this, we used the ddRADseq technique, which generated 1421 SNPs. The heterozygosity values of the analyzed individuals were among the lowest recorded for mammals, ranging from 26 to 91 SNPs/Mb. Furthermore, the individuals from one of the localities, highly isolated due to strong barriers, presented extremely high inbreeding coefficients, with values above 0.7. Under this scenario of low genetic diversity and elevated inbreeding levels, some individuals appeared to be almost genetically identical. We used different methods and simulations to determine if genetic identification and parentage analysis were possible in this population. Only one of the methods, which does not assume population homogeneity, was able to identify all individuals correctly. Therefore, genetically impoverished populations pose a great methodological challenge for their genetic study. However, these populations are of primary scientific and conservation interest, so it is essential to characterize them genetically and improve genomic methodologies for their research.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Using relatedness networks to infer contemporary dispersal: Application to the endangered mammal Galemys pyrenaicus.Mol Ecol. 2017 Jul;26(13):3343-3357. doi: 10.1111/mec.14133. Epub 2017 May 2. Mol Ecol. 2017. PMID: 28374418
-
The genome of the Pyrenean desman and the effects of bottlenecks and inbreeding on the genomic landscape of an endangered species.Evol Appl. 2021 May 29;14(7):1898-1913. doi: 10.1111/eva.13249. eCollection 2021 Jul. Evol Appl. 2021. PMID: 34295371 Free PMC article.
-
Inbreeding and selection shape genomic diversity in captive populations: Implications for the conservation of endangered species.PLoS One. 2017 Apr 19;12(4):e0175996. doi: 10.1371/journal.pone.0175996. eCollection 2017. PLoS One. 2017. PMID: 28423000 Free PMC article.
-
A review of the implications of heterozygosity and inbreeding on germplasm biodiversity and its conservation in the silkworm, Bombyx mori.J Insect Sci. 2011;11:8. doi: 10.1673/031.011.0108. J Insect Sci. 2011. PMID: 21521139 Free PMC article. Review.
-
Reappraisal of the Genetic Diversity Patterns in Puya raimondii-The Queen of the Andes: Insights from Molecular Marker Analysis Reveal an Inbreeding Reproductive Strategy.Plants (Basel). 2025 Jan 22;14(3):321. doi: 10.3390/plants14030321. Plants (Basel). 2025. PMID: 39942885 Free PMC article. Review.
Cited by
-
The Genomics of Isolated Populations of Gampsocleis glabra (Orthoptera: Tettigoniidae) in Central and Western Europe.Insects. 2023 Dec 14;14(12):946. doi: 10.3390/insects14120946. Insects. 2023. PMID: 38132619 Free PMC article.
References
-
- Keller LF, Waller DM. Inbreeding effects in wild populations. Trends Ecol. Evol. 2002;17:230–241. doi: 10.1016/S0169-5347(02)02489-8. - DOI
-
- Frankham, R. et al. Genetic Management of Fragmented Animal and Plant Populations. (Oxford University Press, 2017).
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

