Protein corona: Friend or foe? Co-opting serum proteins for nanoparticle delivery
- PMID: 36503885
- PMCID: PMC9812987
- DOI: 10.1016/j.addr.2022.114635
Protein corona: Friend or foe? Co-opting serum proteins for nanoparticle delivery
Abstract
For systemically delivered nanoparticles to reach target tissues, they must first circulate long enough to reach the target and extravasate there. A challenge is that the particles end up engaging with serum proteins and undergo immune cell recognition and premature clearance. The serum protein binding, also known as protein corona formation, is difficult to prevent, even with artificial protection via "stealth" coating. Protein corona may be problematic as it can interfere with the interaction of targeting ligands with tissue-specific receptors and abrogate the so-called active targeting process, hence, the efficiency of drug delivery. However, recent studies show that serum protein binding to circulating nanoparticles may be actively exploited to enhance their downstream delivery. This review summarizes known issues of protein corona and traditional strategies to control the corona, such as avoiding or overriding its formation, as well as emerging efforts to enhance drug delivery to target organs via nanoparticles. It concludes with a discussion of prevailing challenges in exploiting protein corona for nanoparticle development.
Keywords: Drug delivery; Nanoparticles; Protein corona; Stealth; Targeting.
Copyright © 2022 Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures
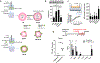




References
-
- Matsumura Y, Maeda H, A New Concept for Macromolecular Therapeutics in Cancer Chemotherapy: Mechanism of Tumoritropic Accumulation of Proteins and the Antitumor Agent Smancs, Cancer Res, 46 (1986) 6387–6392. - PubMed
-
- Singh AK, Chapter 6 - Nanoparticle Pharmacokinetics and Toxicokinetics, in: Singh AK (Ed.) Engineered Nanoparticles, Academic Press, Boston, 2016, pp. 229–293.
-
- Zhang H, Burnum KE, Luna ML, Petritis BO, Kim J-S, Qian W-J, Moore RJ, Heredia-Langner A, Webb-Robertson B-JM, Thrall BD, Camp DG, Smith RD, Pounds JG, Liu T, Quantitative proteomics analysis of adsorbed plasma proteins classifies nanoparticles with different surface properties and size, Proteomics, 11 (2011) 4569–4577. - PMC - PubMed
-
- Monopoli MP, Aberg C, Salvati A, Dawson KA, Biomolecular coronas provide the biological identity of nanosized materials, Nat Nanotechnol, 7 (2012) 779–786. - PubMed
-
- Sepand MR, Ghavami M, Zanganeh S, Stacks S, Ghasemi F, Montazeri H, Corbo C, Derakhshankhah H, Ostad SN, Ghahremani MH, Mahmoudi M, Impact of plasma concentration of transferrin on targeting capacity of nanoparticles, Nanoscale, 12 (2020) 4935–4944. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

