Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
- PMID: 36509735
- PMCID: PMC9744880
- DOI: 10.1038/s41467-022-34970-7
Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Abstract
Extended-spectrum cephalosporins (ESCs) are critically important antimicrobial agents for human and veterinary medicine. ESC resistance (ESC-R) genes have spread worldwide through plasmids and clonal expansion, yet the distribution and dynamics of ESC-R genes in different ecological compartments are poorly understood. Here we use whole genome sequence data of Enterobacterales isolates of human and animal origin from Europe and North America and identify contrasting temporal dynamics. AmpC β-lactamases were initially more dominant in North America in humans and farm animals, only later emerging in Europe. In contrast, specific extended-spectrum β-lactamases (ESBLs) were initially common in animals from Europe and later emerged in North America. This study identifies differences in the relative importance of plasmids and clonal expansion across different compartments for the spread of different ESC-R genes. Understanding the mechanisms of transmission will be critical in the design of interventions to reduce the spread of antimicrobial resistance.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

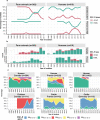
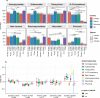

References
-
- Arumugham, V. B. & Cascella, M. Third Generation Cephalosporins. StatPearls (2021). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MR/R000948/1/MRC_/Medical Research Council/United Kingdom
- BBS/E/F/000PR10348/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- CFC-150770/CIHR/Canada
- BB/R012504/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MR/T030062/1/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Medical

