Deletion of the primase-polymerases encoding gene, located in a mobile element in Thermus thermophilus HB27, leads to loss of function mutation of addAB genes
- PMID: 36532486
- PMCID: PMC9751324
- DOI: 10.3389/fmicb.2022.1005862
Deletion of the primase-polymerases encoding gene, located in a mobile element in Thermus thermophilus HB27, leads to loss of function mutation of addAB genes
Abstract
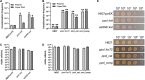
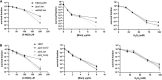
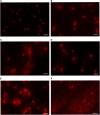
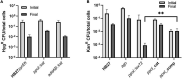
DNA primase-polymerases (Ppol) have been shown to play active roles in DNA repair and damage tolerance, both in prokaryotes and eukaryotes. The ancestral thermophilic bacterium Thermus thermophilus strain HB27 encodes a Ppol protein among the genes present in mobile element ICETh2, absent in other T. thermophilus strains. Using different strategies we ablated the function of Ppol in HB27 cells, either by knocking out the gene through insertional mutagenesis, markerless deletion or through abolition of its catalytic activity. Whole genome sequencing of this diverse collection of Ppol mutants showed spontaneous loss of function mutation in the helicase-nuclease AddAB in every ppol mutant isolated. Given that AddAB is a major player in recombinational repair in many prokaryotes, with similar activity to the proteobacterial RecBCD complex, we have performed a detailed characterization of the ppol mutants in combination with addAB mutants. The results show that knockout addAB mutants are more sensitive to DNA damage agents than the wild type, and present a dramatic three orders of magnitude increase in natural transformation efficiencies with both plasmid and lineal DNA, whereas ppol mutants show defects in plasmid stability. Interestingly, DNA-integrity comet assays showed that the genome of all the ppol and/or addAB mutants was severely affected by widespread fragmentation, however, this did not translate in neat loss of viability of the strains. All these data support that Ppol appears to keep in balance the activity of AddAB as a part of the DNA housekeeping maintenance in T. thermophilus HB27, thus, playing a key role in its genome stability.
Keywords: AddAB; DNA repair; PrimPol; Thermus thermophilus; bacterial transformation; comet assay.
Copyright © 2022 Verdú, Pérez-Arnaiz, Peropadre, Berenguer and Mencía.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

