The UFM1 system regulates ER-phagy through the ufmylation of CYB5R3
- PMID: 36543799
- PMCID: PMC9772183
- DOI: 10.1038/s41467-022-35501-0
The UFM1 system regulates ER-phagy through the ufmylation of CYB5R3
Abstract
Protein modification by ubiquitin-like proteins (UBLs) amplifies limited genome information and regulates diverse cellular processes, including translation, autophagy and antiviral pathways. Ubiquitin-fold modifier 1 (UFM1) is a UBL covalently conjugated with intracellular proteins through ufmylation, a reaction analogous to ubiquitylation. Ufmylation is involved in processes such as endoplasmic reticulum (ER)-associated protein degradation, ribosome-associated protein quality control at the ER and ER-phagy. However, it remains unclear how ufmylation regulates such distinct ER-related functions. Here we identify a UFM1 substrate, NADH-cytochrome b5 reductase 3 (CYB5R3), that localizes on the ER membrane. Ufmylation of CYB5R3 depends on the E3 components UFL1 and UFBP1 on the ER, and converts CYB5R3 into its inactive form. Ufmylated CYB5R3 is recognized by UFBP1 through the UFM1-interacting motif, which plays an important role in the further uyfmylation of CYB5R3. Ufmylated CYB5R3 is degraded in lysosomes, which depends on the autophagy-related protein Atg7- and the autophagy-adaptor protein CDK5RAP3. Mutations of CYB5R3 and genes involved in the UFM1 system cause hereditary developmental disorders, and ufmylation-defective Cyb5r3 knock-in mice exhibit microcephaly. Our results indicate that CYB5R3 ufmylation induces ER-phagy, which is indispensable for brain development.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

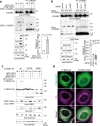




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

