Prostacyclin Released by Cancer-Associated Fibroblasts Promotes Immunosuppressive and Pro-Metastatic Macrophage Polarization in the Ovarian Cancer Microenvironment
- PMID: 36551640
- PMCID: PMC9776493
- DOI: 10.3390/cancers14246154
Prostacyclin Released by Cancer-Associated Fibroblasts Promotes Immunosuppressive and Pro-Metastatic Macrophage Polarization in the Ovarian Cancer Microenvironment
Abstract
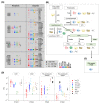
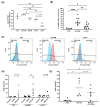
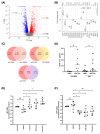
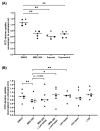
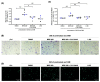
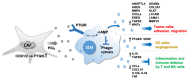
Metastasis of high-grade ovarian carcinoma (HGSC) is orchestrated by soluble mediators of the tumor microenvironment. Here, we have used transcriptomic profiling to identify lipid-mediated signaling pathways encompassing 41 ligand-synthesizing enzymes and 23 cognate receptors in tumor, immune and stroma cells from HGSC metastases and ascites. Due to its strong association with a poor clinical outcome, prostacyclin (PGI2) synthase (PTGIS) is of particular interest in this signaling network. PTGIS is highly expressed by cancer-associated fibroblasts (CAF), concomitant with elevated PGI2 synthesis, whereas tumor-associated macrophages (TAM) exhibit the highest expression of its surface receptor (PTGIR). PTGIR activation by PGI2 agonists triggered cAMP accumulation and induced a mixed-polarization macrophage phenotype with altered inflammatory gene expression, including CXCL10 and IL12A repression, as well as reduced phagocytic capability. Co-culture experiments provided further evidence for the interaction of CAF with macrophages via PGI2, as the effect of PGI2 agonists on phagocytosis was mitigated by cyclooxygenase inhibitors. Furthermore, conditioned medium from PGI2-agonist-treated TAM promoted tumor adhesion to mesothelial cells and migration in a PTGIR-dependent manner, and PTGIR activation induced the expression of metastasis-associated and pro-angiogenic genes. Taken together, our study identifies a PGI2/PTGIR-driven crosstalk between CAF, TAM and tumor cells, promoting immune suppression and a pro-metastatic environment.
Keywords: carcinoma-associated fibroblasts; high-grade serous ovarian carcinoma; pro-metastatic phenotype; prostacyclin; signaling network; tumor-associated macrophages.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
The multicellular signalling network of ovarian cancer metastases.Clin Transl Med. 2021 Nov;11(11):e633. doi: 10.1002/ctm2.633. Clin Transl Med. 2021. PMID: 34841720 Free PMC article.
-
Hypoxia-hindered methylation of PTGIS in endometrial stromal cells accelerates endometriosis progression by inducing CD16- NK-cell differentiation.Exp Mol Med. 2022 Jul;54(7):890-905. doi: 10.1038/s12276-022-00793-1. Epub 2022 Jul 4. Exp Mol Med. 2022. PMID: 35781537 Free PMC article.
-
Prostacyclin receptor (PTGIR) in the porcine endometrium: Regulation of expression and role in luminal epithelial and stromal cells.Theriogenology. 2015 Oct 1;84(6):969-82. doi: 10.1016/j.theriogenology.2015.05.034. Epub 2015 Jun 6. Theriogenology. 2015. PMID: 26139576
-
Tumor-associated macrophages promote ovarian cancer cell migration by secreting transforming growth factor beta induced (TGFBI) and tenascin C.Cell Death Dis. 2020 Apr 20;11(4):249. doi: 10.1038/s41419-020-2438-8. Cell Death Dis. 2020. PMID: 32312959 Free PMC article.
-
Targeting Tyro3, Axl and MerTK (TAM receptors): implications for macrophages in the tumor microenvironment.Mol Cancer. 2019 May 14;18(1):94. doi: 10.1186/s12943-019-1022-2. Mol Cancer. 2019. PMID: 31088471 Free PMC article. Review.
Cited by
-
Macrophage heterogeneity and its interactions with stromal cells in tumour microenvironment.Cell Biosci. 2024 Feb 1;14(1):16. doi: 10.1186/s13578-024-01201-z. Cell Biosci. 2024. PMID: 38303024 Free PMC article. Review.
-
Identification of a cancer associated fibroblasts-related index to predict prognosis and immune landscape in ovarian cancer.Sci Rep. 2023 Dec 7;13(1):21565. doi: 10.1038/s41598-023-48653-w. Sci Rep. 2023. PMID: 38057405 Free PMC article.
-
The impact of the tumor microenvironment on macrophages.Front Immunol. 2025 May 16;16:1572764. doi: 10.3389/fimmu.2025.1572764. eCollection 2025. Front Immunol. 2025. PMID: 40453088 Free PMC article. Review.
-
Arachidonic acid metabolism in health and disease.MedComm (2020). 2023 Sep 20;4(5):e363. doi: 10.1002/mco2.363. eCollection 2023 Oct. MedComm (2020). 2023. PMID: 37746665 Free PMC article. Review.
-
Knockdown TNF family prognosis index crucial gene PDE4B promoted PANoptosis of ovarian carcinoma cell:Based in vitro and in vivo experiments.Transl Oncol. 2025 Jun;56:102333. doi: 10.1016/j.tranon.2025.102333. Epub 2025 Apr 16. Transl Oncol. 2025. PMID: 40245751 Free PMC article.
References
-
- Reinartz S., Lieber S., Pesek J., Brandt D.T., Asafova A., Finkernagel F., Watzer B., Nockher W.A., Nist A., Stiewe T., et al. Cell type-selective pathways and clinical associations of lysophosphatidic acid biosynthesis and signaling in the ovarian cancer microenvironment. Mol. Oncol. 2019;13:185–201. doi: 10.1002/1878-0261.12396. - DOI - PMC - PubMed
-
- Reinartz S., Finkernagel F., Adhikary T., Rohnalter V., Schumann T., Schober Y., Nockher W.A., Nist A., Stiewe T., Jansen J.M., et al. A transcriptome-based global map of signaling pathways in the ovarian cancer microenvironment associated with clinical outcome. Genome Biol. 2016;17:108. doi: 10.1186/s13059-016-0956-6. - DOI - PMC - PubMed
-
- Dietze R., Hammoud M.K., Gómez-Serrano M., Unger A., Bieringer T., Finkernagel F., Sokol A.M., Nist A., Stiewe T., Reinartz S., et al. Phosphoproteomics identify arachidonic-acid-regulated signal transduction pathways modulating macrophage functions with implications for ovarian cancer. Theranostics. 2021;11:1377–1395. doi: 10.7150/thno.52442. - DOI - PMC - PubMed
-
- Hammoud M.K., Dietze R., Pesek J., Finkernagel F., Unger A., Bieringer T., Nist A., Stiewe T., Bhagwat A.M., Nockher W.A., et al. Arachidonic acid, a clinically adverse mediator in the ovarian cancer microenvironment, impairs JAK-STAT signaling in macrophages by perturbing lipid raft structures. Mol. Oncol. 2022;16:3146–3166. doi: 10.1002/1878-0261.13221. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

