Comparative RNA-Sequencing Analysis Reveals High Complexity and Heterogeneity of Transcriptomic and Immune Profiles in Hepatocellular Carcinoma Tumors of Viral (HBV, HCV) and Non-Viral Etiology
- PMID: 36557005
- PMCID: PMC9785216
- DOI: 10.3390/medicina58121803
Comparative RNA-Sequencing Analysis Reveals High Complexity and Heterogeneity of Transcriptomic and Immune Profiles in Hepatocellular Carcinoma Tumors of Viral (HBV, HCV) and Non-Viral Etiology
Abstract
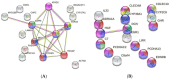
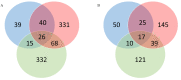
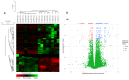
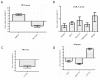
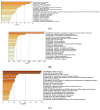
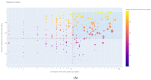
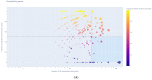
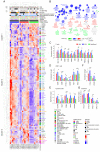
Background and Objectives: Hepatocellular carcinoma (HCC), the most common type of primary liver cancer, is the leading cause of cancer-related mortality. It arises and progresses against fibrotic or cirrhotic backgrounds mainly due to infection with hepatitis viruses B (HBV) or C (HCV) or non-viral causes that lead to chronic inflammation and genomic changes. A better understanding of molecular and immune mechanisms in HCC subtypes is needed. Materials and Methods: To identify transcriptional changes in primary HCC tumors with or without hepatitis viral etiology, we analyzed the transcriptomes of 24 patients by next-generation sequencing. Results: We identified common and unique differentially expressed genes for each etiological tumor group and analyzed the expression of SLC, ATP binding cassette, cytochrome 450, cancer testis, and heat shock protein genes. Metascape functional enrichment analysis showed mainly upregulated cell-cycle pathways in HBV and HCV and upregulated cell response to stress in non-viral infection. GeneWalk analysis identified regulator, hub, and moonlighting genes and highlighted CCNB1, ACTN2, BRCA1, IGF1, CDK1, AURKA, AURKB, and TOP2A in the HCV group and HSF1, HSPA1A, HSP90AA1, HSPB1, HSPA5, PTK2, and AURKB in the group without viral infection as hub genes. Immune infiltrate analysis showed that T cell, cytotoxic, and natural killer cell markers were significantly more highly expressed in HCV than in non-viral tumors. Genes associated with monocyte activation had the highest expression levels in HBV, while high expression of genes involved in primary adaptive immune response and complement receptor activity characterized tumors without viral infection. Conclusions: Our comprehensive study underlines the high degree of complexity of immune profiles in the analyzed groups, which adds to the heterogeneous HCC genomic landscape. The biomarkers identified in each HCC group might serve as therapeutic targets.
Keywords: HBV; HCV; RNA-seq; biomarkers; hepatocellular carcinoma; immune infiltrate.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
-
- Globocan. [(accessed on 1 May 2021)]. Available online: https://gco.iarc.fr/today.
-
- Bruix J., Qin S., Merle P., Granito A., Huang Y.H., Bodoky G., Pracht M., Yokosuka O., Rosmorduc O., Breder V., et al. Regorafenib for patients with hepatocellular carcinoma who progressed on sorafenib treatment (RESORCE): A randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;389:56–66. doi: 10.1016/S0140-6736(16)32453-9. - DOI - PubMed
-
- Zhu A.X., Park J.O., Ryoo B.Y., Yen C.J., Poon R., Pastorelli D., Blanc J.F., Chung H.C., Baron A.D., Pfiffer T.E., et al. Ramucirumab versus placebo as second-line treatment in patients with advanced hepatocellular carcinoma following first-line therapy with sorafenib (REACH): A randomised, double-blind, multicentre, phase 3 trial. Lancet Oncol. 2015;16:859–870. doi: 10.1016/S1470-2045(15)00050-9. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

