Genome-Wide Identification and Characterization of TCP Gene Family Members in Melastoma candidum
- PMID: 36558169
- PMCID: PMC9787641
- DOI: 10.3390/molecules27249036
Genome-Wide Identification and Characterization of TCP Gene Family Members in Melastoma candidum
Abstract
It has been confirmed that the plant-specific Teosinte-branched 1/Cycloidea/Proliferating (TCP) gene family plays a pivotal role during plant growth and development. M. candidum is a native ornamental species and has a wide range of pharmacodynamic effects. However, there is still a lack of research on TCP’s role in controlling M. candidum’s development, abiotic stress responses and hormone metabolism. A comprehensive description of the TCP gene family in M. candidum is urgently needed. In this study, we used the HMMER search method in conjunction with the BLASTp method to identify the members of the TCP gene family, and a total of 35 TCP genes were identified. A domain analysis further confirmed that all 35 TCPs contained a TCP superfamily, a characteristic involved in dimerization and DNA binding that can be found in most genes from this gene family, suggesting that our identification was effective. As a result of the domain conservation analysis, the 35 TCP genes could be classified into two classes, TCP-P and TCP-C, based on the conservative regions of 55 and 59 amino acids, respectively. Gene-duplication analysis revealed that most TCP genes were present in duplication events that eventually led to TCP gene expansion in M. candidum. All the detected gene pairs had a Ka/Ks value of less than one, suggesting that purification selection is the most important factor that influences the evolution of TCP genes. Phylogenetic analysis of three species displayed the evolutionary relationship of TCP genes across different species and further confirmed our results. The real-time quantitative PCR (qRT-PCR) results showed that McTCP2a, McTCP7a, McTCP10, McTCP11, McTCP12a, McTCP13, McTCP16, McTCP17, McTCP18, McTCP20 and McTCP21 may be involved in leaf development; McTCP4a, McTCP1, McTCP14, McTCP17, McTCP18, McTCP20, McTCP22 and McTCP24 may be involved in flower development; and McTCP2a, McTCP3, McTCP5a, McTCP6, McTCP7a, McTCP9, McTCP11, McTCP14 and McTCP16 may be involved in seed development. Our results dissect the TCP gene family across the genome of M. candidum and provide valuable information for exploring TCP genes to promote molecular breeding and property improvement of M. candidum in the future.
Keywords: Melastoma candidum; TCP gene family; collinearity analysis; gene pairs; phylogenetic analysis.
Conflict of interest statement
The authors declare no conflict of interest.
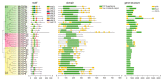
Figures









Similar articles
-
Genome-Wide Identification and Expression Analysis of TCP Transcription Factors Responding to Multiple Stresses in Arachis hypogaea L.Int J Mol Sci. 2025 Jan 26;26(3):1069. doi: 10.3390/ijms26031069. Int J Mol Sci. 2025. PMID: 39940846 Free PMC article.
-
Genome-wide identification and expression profiling analysis of maize AP2/ERF superfamily genes reveal essential roles in abiotic stress tolerance.BMC Genomics. 2022 Feb 12;23(1):125. doi: 10.1186/s12864-022-08345-7. BMC Genomics. 2022. PMID: 35151253 Free PMC article.
-
Genome-Wide Identification and Characterization of TCP Genes in Eight Prunus Species and Their Expression Patterns Under Cold Stress in P. tenella var. tenella.Genes (Basel). 2024 Nov 8;15(11):1443. doi: 10.3390/genes15111443. Genes (Basel). 2024. PMID: 39596643 Free PMC article.
-
TCP Transcription Factors in Plant Reproductive Development: Juggling Multiple Roles.Biomolecules. 2023 Apr 26;13(5):750. doi: 10.3390/biom13050750. Biomolecules. 2023. PMID: 37238620 Free PMC article. Review.
-
The Arabidopsis thaliana TCP transcription factors: A broadening horizon beyond development.Plant Signal Behav. 2015;10(7):e1044192. doi: 10.1080/15592324.2015.1044192. Plant Signal Behav. 2015. PMID: 26039357 Free PMC article. Review.
Cited by
-
Antifungal Action of Arabidopsis thaliana TCP21 via Induction of Oxidative Stress and Apoptosis.Antioxidants (Basel). 2023 Sep 15;12(9):1767. doi: 10.3390/antiox12091767. Antioxidants (Basel). 2023. PMID: 37760070 Free PMC article.
-
Genome-wide identification and expression analysis of TCP family genes in Catharanthus roseus.Front Plant Sci. 2023 Apr 12;14:1161534. doi: 10.3389/fpls.2023.1161534. eCollection 2023. Front Plant Sci. 2023. PMID: 37123846 Free PMC article.
-
Genome-wide identification of TCP transcription factors and their potential roles in hydrolyzable tannin production in Quercus variabilis cupule.Front Plant Sci. 2024 Aug 6;15:1444081. doi: 10.3389/fpls.2024.1444081. eCollection 2024. Front Plant Sci. 2024. PMID: 39166255 Free PMC article.
-
Integrating genome and transcriptome-wide data to explore the expression dynamics of TCP genes in Pisum sativum under salt stress.Front Plant Sci. 2025 May 1;16:1580890. doi: 10.3389/fpls.2025.1580890. eCollection 2025. Front Plant Sci. 2025. PMID: 40385237 Free PMC article.
-
Genome-Wide Identification and Analysis of TCP Gene Family among Three Dendrobium Species.Plants (Basel). 2023 Sep 7;12(18):3201. doi: 10.3390/plants12183201. Plants (Basel). 2023. PMID: 37765364 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

