An epithelial-immune circuit amplifies inflammasome and IL-6 responses to SARS-CoV-2
- PMID: 36563691
- PMCID: PMC9731922
- DOI: 10.1016/j.chom.2022.12.005
An epithelial-immune circuit amplifies inflammasome and IL-6 responses to SARS-CoV-2
Abstract
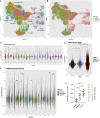
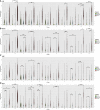
Elevated levels of cytokines IL-1β and IL-6 are associated with severe COVID-19. Investigating the underlying mechanisms, we find that while primary human airway epithelia (HAE) have functional inflammasomes and support SARS-CoV-2 replication, they are not the source of IL-1β released upon infection. In leukocytes, the SARS-CoV-2 E protein upregulates inflammasome gene transcription via TLR2 to prime, but not activate, inflammasomes. SARS-CoV-2-infected HAE supply a second signal, which includes genomic and mitochondrial DNA, to stimulate leukocyte IL-1β release. Nuclease treatment, STING, and caspase-1 inhibition but not NLRP3 inhibition blocked leukocyte IL-1β release. After release, IL-1β stimulates IL-6 secretion from HAE. Therefore, infection alone does not increase IL-1β secretion by either cell type. Rather, bi-directional interactions between the SARS-CoV-2-infected epithelium and immune bystanders stimulates both IL-1β and IL-6, creating a pro-inflammatory cytokine circuit. Consistent with these observations, patient autopsy lungs show elevated myeloid inflammasome gene signatures in severe COVID-19.
Keywords: AIM2; IL-1β; IL-6; NLRP1; NLRP3; SARS-CoV-2; STING; TLR2; inflammasome.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests J.P.-Y.T. is a cofounder of IMMvention Therapeutix, which works on inflammasome inhibitors.
Figures





Comment in
-
Stoking inflammasome fires in the COVID-19 neighborhood.Cell Host Microbe. 2023 Feb 8;31(2):168-170. doi: 10.1016/j.chom.2023.01.008. Cell Host Microbe. 2023. PMID: 36758516 Free PMC article.
References
-
- Zhou F., Yu T., Du R., Fan G., Liu Y., Liu Z., Xiang J., Wang Y., Song B., Gu X., et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395:1054–1062. doi: 10.1016/S0140-6736(20)30566-3. - DOI - PMC - PubMed
-
- Wilk A.J., Rustagi A., Zhao N.Q., Roque J., Martínez-Colón G.J., McKechnie J.L., Ivison G.T., Ranganath T., Vergara R., Hollis T., et al. A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat. Med. 2020;26:1070–1076. doi: 10.1038/s41591-020-0944-y. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
- R01 HL136961/HL/NHLBI NIH HHS/United States
- R01 AI157253/AI/NIAID NIH HHS/United States
- R01 AI158314/AI/NIAID NIH HHS/United States
- R01 ES032643/ES/NIEHS NIH HHS/United States
- R56 AI029564/AI/NIAID NIH HHS/United States
- R01 DE031951/DE/NIDCR NIH HHS/United States
- T32 CA071341/CA/NCI NIH HHS/United States
- R01 AI029564/AI/NIAID NIH HHS/United States
- UH3 HL123645/HL/NHLBI NIH HHS/United States
- R01 DE026728/DE/NIDCR NIH HHS/United States
- R37 AI029564/AI/NIAID NIH HHS/United States
- U01 DE029255/DE/NIDCR NIH HHS/United States
- UC6 AI058607/AI/NIAID NIH HHS/United States
- P01 HL108808/HL/NHLBI NIH HHS/United States
- U19 AI100625/AI/NIAID NIH HHS/United States
- R56 AI158314/AI/NIAID NIH HHS/United States
- R35 CA232109/CA/NCI NIH HHS/United States
- R01 AI141333/AI/NIAID NIH HHS/United States
- P30 DK065988/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

