Rapamycin improves Graves' orbitopathy by suppressing CD4+ cytotoxic T lymphocytes
- PMID: 36580373
- PMCID: PMC9977423
- DOI: 10.1172/jci.insight.160377
Rapamycin improves Graves' orbitopathy by suppressing CD4+ cytotoxic T lymphocytes
Abstract
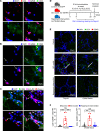
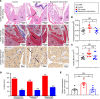
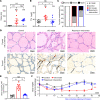
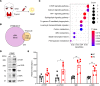
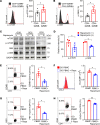
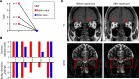
CD4+ cytotoxic T lymphocytes (CTLs) were recently implicated in immune-mediated inflammation and fibrosis progression of Graves' orbitopathy (GO). However, little is known about therapeutic targeting of CD4+ CTLs. Herein, we studied the effect of rapamycin, an approved mTOR complex 1 (mTORC1) inhibitor, in a GO mouse model, in vitro, and in patients with refractory GO. In the adenovirus-induced model, rapamycin significantly decreased the incidence of GO. This was accompanied by the reduction of both CD4+ CTLs and the reduction of orbital inflammation, adipogenesis, and fibrosis. CD4+ CTLs from patients with active GO showed upregulation of the mTOR pathway, while rapamycin decreased their proportions and cytotoxic function. Low-dose rapamycin treatment substantially improved diplopia and the clinical activity score in steroid-refractory patients with GO. Single-cell RNA-Seq revealed that eye motility improvement was closely related to suppression of inflammation and chemotaxis in CD4+ CTLs. In conclusion, rapamycin is a promising treatment for CD4+ CTL-mediated inflammation and fibrosis in GO.
Keywords: Autoimmune diseases; Autoimmunity; Endocrinology; Immunotherapy; T cells.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

