Tissue fluidification promotes a cGAS-STING cytosolic DNA response in invasive breast cancer
- PMID: 36581770
- PMCID: PMC10156599
- DOI: 10.1038/s41563-022-01431-x
Tissue fluidification promotes a cGAS-STING cytosolic DNA response in invasive breast cancer
Erratum in
-
Author Correction: Tissue fluidification promotes a cGAS-STING cytosolic DNA response in invasive breast cancer.Nat Mater. 2023 Mar;22(3):400. doi: 10.1038/s41563-023-01479-3. Nat Mater. 2023. PMID: 36702890 Free PMC article. No abstract available.
Abstract
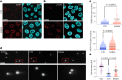
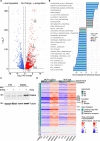
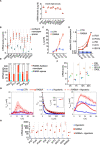
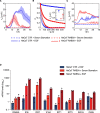
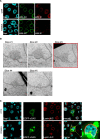
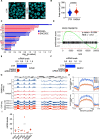
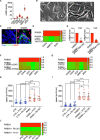
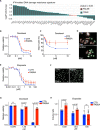
The process in which locally confined epithelial malignancies progressively evolve into invasive cancers is often promoted by unjamming, a phase transition from a solid-like to a liquid-like state, which occurs in various tissues. Whether this tissue-level mechanical transition impacts phenotypes during carcinoma progression remains unclear. Here we report that the large fluctuations in cell density that accompany unjamming result in repeated mechanical deformations of cells and nuclei. This triggers a cellular mechano-protective mechanism involving an increase in nuclear size and rigidity, heterochromatin redistribution and remodelling of the perinuclear actin architecture into actin rings. The chronic strains and stresses associated with unjamming together with the reduction of Lamin B1 levels eventually result in DNA damage and nuclear envelope ruptures, with the release of cytosolic DNA that activates a cGAS-STING (cyclic GMP-AMP synthase-signalling adaptor stimulator of interferon genes)-dependent cytosolic DNA response gene program. This mechanically driven transcriptional rewiring ultimately alters the cell state, with the emergence of malignant traits, including epithelial-to-mesenchymal plasticity phenotypes and chemoresistance in invasive breast carcinoma.
© 2022. The Author(s).
Conflict of interest statement
M.P. is co-founder, member of the board of directors and stakeholder of the company CheckmAb s.r.l. The remaining authors declare no competing interests.
Figures








Comment in
-
Unjamming tumour cell invasion through cGAS-STING.Nat Mater. 2023 May;22(5):532-533. doi: 10.1038/s41563-023-01542-z. Nat Mater. 2023. PMID: 37138010 No abstract available.
References
-
- Northey JJ, Przybyla L, Weaver VM. Tissue force programs cell fate and tumor aggression. Cancer Discov. 2017;7:1224–1237. doi: 10.1158/2159-8290.CD-16-0733. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

