Cost-saving population genomic investigation of Daphnia longispina complex resting eggs using whole-genome amplification and pre-sequencing screening
- PMID: 36582775
- PMCID: PMC9793289
- DOI: 10.1002/ece3.9682
Cost-saving population genomic investigation of Daphnia longispina complex resting eggs using whole-genome amplification and pre-sequencing screening
Abstract
Resting stages of aquatic organisms that accumulate in the sediment over time are an exceptional resource that allows direct insights into past populations and addressing evolutionary questions. This is of particular interest in taxa that face relatively new environmental challenges, e.g., climate change and eutrophication, such as the Daphnia longispina species complex, a keystone zooplankton group in European freshwater ecosystems. However, genomic analysis might be challenging as DNA yield from many of these resting stages can be low and the material degraded. To reliably allow the resequencing of single Daphnia resting eggs from different sediment layers and characterize genomic changes through time, we performed whole-genome amplification to obtain DNA amounts suitable for genome resequencing and tested multiple protocols involving egg isolation, whole-genome amplification kits, and library preparation. A pre-sequencing contamination screening was developed, consisting of amplifying mitochondrial Daphnia and bacterial markers, to quickly assess and exclude possibly contaminated samples. In total, we successfully amplified and sequenced nine genomes from Daphnia resting eggs that could be identified as Daphnia longispina species. We analyzed the genome coverage and heterozygosity of these samples to optimize this method for future projects involving population genomic investigation of the resting egg bank.
Keywords: Daphnia longispina species complex; population genomics; resting egg bank; whole‐genome amplification.
© 2022 The Authors. Ecology and Evolution published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
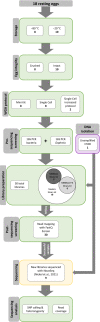

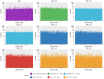
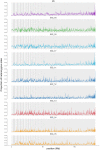
Similar articles
-
Whole genome amplification and sequencing of a Daphnia resting egg.Mol Ecol Resour. 2018 Jan;18(1):118-127. doi: 10.1111/1755-0998.12720. Epub 2017 Oct 14. Mol Ecol Resour. 2018. PMID: 28926213
-
Hybridization Dynamics and Extensive Introgression in the Daphnia longispina Species Complex: New Insights from a High-Quality Daphnia galeata Reference Genome.Genome Biol Evol. 2021 Dec 1;13(12):evab267. doi: 10.1093/gbe/evab267. Genome Biol Evol. 2021. PMID: 34865004 Free PMC article.
-
Refining the evolutionary time machine: An assessment of whole genome amplification using single historical Daphnia eggs.Mol Ecol Resour. 2022 Apr;22(3):946-961. doi: 10.1111/1755-0998.13524. Epub 2021 Oct 20. Mol Ecol Resour. 2022. PMID: 34672105
-
Optimized and affordable high-throughput sequencing workflow for preserved and nonpreserved small zooplankton specimens.Mol Ecol Resour. 2020 Nov;20(6):1632-1646. doi: 10.1111/1755-0998.13228. Epub 2020 Aug 9. Mol Ecol Resour. 2020. PMID: 32677266
-
An integrated multi-disciplinary approach for studying multiple stressors in freshwater ecosystems: Daphnia as a model organism.Integr Comp Biol. 2011 Oct;51(4):623-33. doi: 10.1093/icb/icr103. Epub 2011 Aug 27. Integr Comp Biol. 2011. PMID: 21873644 Review.
References
-
- Andrews, S. (2010). FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
-
- Blattner, L. , Lucek, K. , Beck, N. , Berner, D. , & Von Fumetti, S. (2022). Intra‐Alpine Islands: Population genomic inference reveals high degree of isolation between freshwater spring habitats. Diversity and Distributions, 28(2), 291–305. 10.1111/ddi.13461 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

