Pathogenesis-related protein 1 suppresses oomycete pathogen by targeting against AMPK kinase complex
- PMID: 36585103
- PMCID: PMC9811325
- DOI: 10.1016/j.jare.2022.02.002
Pathogenesis-related protein 1 suppresses oomycete pathogen by targeting against AMPK kinase complex
Abstract
Introduction: During the arms race between plants and pathogens, pathogenesis-related proteins (PR) in host plants play a crucial role in disease resistance, especially PR1. PR1 constitute a secretory peptide family, and their role in plant defense has been widely demonstrated in both hosts and in vitro. However, the mechanisms by which they control host-pathogen interactions and the nature of their targets within the pathogen remain poorly understood.
Objectives: The present study was aimed to investigate the anti-oomycete activity of secretory PR1 proteins and elaborate their underlying mechanisms.
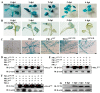
Methods: This study was conducted in the potato-Phytophthora infestans pathosystem. After being induced by the pathogen infection, the cross-kingdom translocation of secretory PR1 was demonstrated by histochemical assays and western blot, and their targets in P. infestans were identified by yeast-two-hybrid assays, bimolecular fluorescence complementation assays, and co-immunoprecipitation assay.
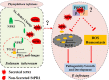
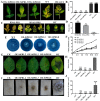
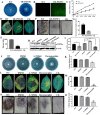
Results: The results showed that the expression of secretory PR1-encoding genes was induced during pathogen infection, and the host could deliver PR1 into P. infestans to inhibit its vegetative growth and pathogenicity. The translocated secretory PR1 targeted the subunits of the AMPK kinase complex in P. infestans, thus affecting the AMPK-driven phosphorylation of downstream target proteins, preventing ROS homeostasis, and down-regulating the expression of RxLR effectors.
Conclusion: The results provide novel insights into the molecular function of PR1 in protecting plants against pathogen infection, and uncover a potential target for preventing pre- and post-harvest late blight.
Keywords: AMPK kinase complex; Cross-kingdom translocation; Host-pathogen interaction; Pathogenesis-related protein 1; Phytophthora infestans; Potato late blight.
Copyright © 2022. Production and hosting by Elsevier B.V.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures





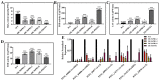
Similar articles
-
The Phytophthora infestans effector Pi05910 suppresses and destabilizes host glycolate oxidase StGOX4 to promote plant susceptibility.Mol Plant Pathol. 2024 Nov;25(11):e70021. doi: 10.1111/mpp.70021. Mol Plant Pathol. 2024. PMID: 39487604 Free PMC article.
-
Metabolic Model of the Phytophthora infestans-Tomato Interaction Reveals Metabolic Switches during Host Colonization.mBio. 2019 Jul 9;10(4):e00454-19. doi: 10.1128/mBio.00454-19. mBio. 2019. PMID: 31289172 Free PMC article.
-
Phytophthora infestans RXLR effector Pi23014 targets host RNA-binding protein NbRBP3a to suppress plant immunity.Mol Plant Pathol. 2024 Jan;25(1):e13416. doi: 10.1111/mpp.13416. Mol Plant Pathol. 2024. PMID: 38279850 Free PMC article.
-
Pathogen virulence of Phytophthora infestans: from gene to functional genomics.Physiol Mol Biol Plants. 2013 Apr;19(2):165-77. doi: 10.1007/s12298-012-0157-z. Physiol Mol Biol Plants. 2013. Retraction in: Physiol Mol Biol Plants. 2015 Jan;21(1):167. doi: 10.1007/s12298-014-0263-1. PMID: 24431484 Free PMC article. Retracted. Review.
-
Late blight in tomato: insights into the pathogenesis of the aggressive pathogen Phytophthora infestans and future research priorities.Planta. 2021 May 8;253(6):119. doi: 10.1007/s00425-021-03636-x. Planta. 2021. PMID: 33963935 Review.
Cited by
-
Cotton STARD Gene Family: Characterization, Evolution, and Expression Profiles During Salt Stress.Genes (Basel). 2025 Jul 11;16(7):813. doi: 10.3390/genes16070813. Genes (Basel). 2025. PMID: 40725469 Free PMC article.
-
Identification of the Function of the Pathogenesis-Related Protein GmPR1L in the Resistance of Soybean to Cercospora sojina Hara.Genes (Basel). 2023 Apr 15;14(4):920. doi: 10.3390/genes14040920. Genes (Basel). 2023. PMID: 37107678 Free PMC article.
-
Validation of Monilinia fructicola Putative Effector Genes in Different Host Peach (Prunus persica) Cultivars and Defense Response Investigation.J Fungi (Basel). 2025 Jan 6;11(1):39. doi: 10.3390/jof11010039. J Fungi (Basel). 2025. PMID: 39852458 Free PMC article.
-
Mechanisms of Chinese Hickory Resistance to Dry Rot Disease by Botryosphaeria dothidea: A Comprehensive Analysis from Gene Expression to Non-Coding RNAs.Plants (Basel). 2025 Mar 4;14(5):793. doi: 10.3390/plants14050793. Plants (Basel). 2025. PMID: 40094748 Free PMC article.
-
The Phytophthora sojae nuclear effector PsAvh110 targets a host transcriptional complex to modulate plant immunity.Plant Cell. 2023 Jan 2;35(1):574-597. doi: 10.1093/plcell/koac300. Plant Cell. 2023. PMID: 36222564 Free PMC article.
References
-
- Haverkort A.J., Struik P.C., Visser R.G.F., Jacobsen E. Applied biotechnology to combat late blight in potato caused by Phytophthora infestans. Potato Res. 2009;52(3):249–264.
-
- Dong X. NPR1, all things considered. Curr Opin Plant Biol. 2004;7(5):547–552. - PubMed
-
- Van Loon L.C., Van Kammen A. Polyacrylamide disc electro-phoresis of the soluble leaf proteins from Nicotiana tabacum var. 'Samsun' and 'Samsun NN'. II. Changes in protein constitution after infection with tobacco mosaic virus. Virology. 1970;40(2):199–211. - PubMed
-
- van Loon L.C., Rep M., Pieterse C.M.J. Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol. 2006;44(1):135–162. - PubMed
-
- Niderman T., Genetet I., Bruyere T., Gees R., Stintzi A., Legrand M., et al. Pathogenesis-Related PR-1 proteins are antifungal (isolation and characterization of three 14-kilodalton proteins of tomato and of a basic PR-1 of tobacco with inhibitory activity against Phytophthora infestans) Plant Physiol. 1995;108(1):17–27. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

