Targeting T cell checkpoints 41BB and LAG3 and myeloid cell CXCR1/CXCR2 results in antitumor immunity and durable response in pancreatic cancer
- PMID: 36585453
- PMCID: PMC9925045
- DOI: 10.1038/s43018-022-00500-z
Targeting T cell checkpoints 41BB and LAG3 and myeloid cell CXCR1/CXCR2 results in antitumor immunity and durable response in pancreatic cancer
Abstract
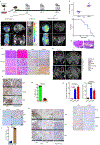
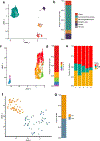
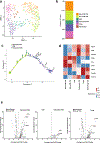
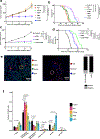
Pancreatic ductal adenocarcinoma (PDAC) is considered non-immunogenic, with trials showing its recalcitrance to PD1 and CTLA4 immune checkpoint therapies (ICTs). Here, we sought to systematically characterize the mechanisms underlying de novo ICT resistance and to identify effective therapeutic options for PDAC. We report that agonist 41BB and antagonist LAG3 ICT alone and in combination, increased survival and antitumor immunity, characterized by modulating T cell subsets with antitumor activity, increased T cell clonality and diversification, decreased immunosuppressive myeloid cells and increased antigen presentation/decreased immunosuppressive capability of myeloid cells. Translational analyses confirmed the expression of 41BB and LAG3 in human PDAC. Since single and dual ICTs were not curative, T cell-activating ICTs were combined with a CXCR1/2 inhibitor targeting immunosuppressive myeloid cells. Triple therapy resulted in durable complete responses. Given similar profiles in human PDAC and the availability of these agents for clinical testing, our findings provide a testable hypothesis for this lethal disease.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Declaration of Interests
R.A.D. is Founder, Advisor and/or Director of Tvardi Therapeutics, Asylia Therapeutics, Stellanova Therapeutics, Nirogy Therapeutics, and Sporos Bioventures. J.A.Z. is the President and CEO, D.Y.M. is the Director of Medicinal Chemistry and Preclinical Development at Syntrix Pharmaceuticals. A.M. receives royalties from Cosmos Wisdom Biotechnology and Thrive Earlier Detection, an Exact Sciences Company. A.M. is also a consultant for Freenome and Tezcat Biotechnology. The other authors declare no competing interests.
Figures














Comment in
-
Targeting T cell checkpoints and myeloid suppressor cells is effective in pancreatic cancer.Nat Cancer. 2023 Jan;4(1):7-8. doi: 10.1038/s43018-022-00501-y. Nat Cancer. 2023. PMID: 36639509 No abstract available.
References
-
- Siegel RL, Miller KD, Fuchs HE, and Jemal A. Cancer statistics, 2022. CA: Cancer Journal for Clinicians 2022; 72(1):7–33. - PubMed
-
- Rahib L, Smith BD, Aizenberg R, et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res 2014; 74(11):2913–2921. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

