A universal sequencing read interpreter
- PMID: 36598975
- PMCID: PMC9812397
- DOI: 10.1126/sciadv.add2793
A universal sequencing read interpreter
Abstract
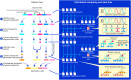
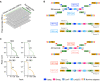
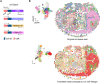
Massively parallel DNA sequencing has led to the rapid growth of highly multiplexed experiments in biology. These experiments produce unique sequencing results that require specific analysis pipelines to decode highly structured reads. However, no versatile framework that interprets sequencing reads to extract their encoded information for downstream biological analysis has been developed. Here, we report INTERSTELLAR (interpretation, scalable transformation, and emulation of large-scale sequencing reads) that decodes data values encoded in theoretically any type of sequencing read and translates them into sequencing reads of another structure of choice. We demonstrated that INTERSTELLAR successfully extracted information from a range of short- and long-read sequencing reads and translated those of single-cell (sc)RNA-seq, scATAC-seq, and spatial transcriptomics to be analyzed by different software tools that have been developed for conceptually the same types of experiments. INTERSTELLAR will greatly facilitate the development of sequencing-based experiments and sharing of data analysis pipelines.
Figures




Similar articles
-
Split Pool Ligation-based Single-cell Transcriptome sequencing (SPLiT-seq) data processing pipeline comparison.BMC Genomics. 2024 Apr 12;25(1):361. doi: 10.1186/s12864-024-10285-3. BMC Genomics. 2024. PMID: 38609853 Free PMC article.
-
Apollo: a sequencing-technology-independent, scalable and accurate assembly polishing algorithm.Bioinformatics. 2020 Jun 1;36(12):3669-3679. doi: 10.1093/bioinformatics/btaa179. Bioinformatics. 2020. PMID: 32167530
-
Querying large read collections in main memory: a versatile data structure.BMC Bioinformatics. 2011 Jun 17;12:242. doi: 10.1186/1471-2105-12-242. BMC Bioinformatics. 2011. PMID: 21682852 Free PMC article.
-
A comprehensive evaluation of long read error correction methods.BMC Genomics. 2020 Dec 21;21(Suppl 6):889. doi: 10.1186/s12864-020-07227-0. BMC Genomics. 2020. PMID: 33349243 Free PMC article. Review.
-
Mapping RNA-seq reads to transcriptomes efficiently based on learning to hash method.Comput Biol Med. 2020 Jan;116:103539. doi: 10.1016/j.compbiomed.2019.103539. Epub 2019 Nov 13. Comput Biol Med. 2020. PMID: 31765913 Review.
Cited by
-
A multi-kingdom genetic barcoding system for precise clone isolation.Nat Biotechnol. 2025 May 21. doi: 10.1038/s41587-025-02649-1. Online ahead of print. Nat Biotechnol. 2025. PMID: 40399693
-
Applications of single‑cell omics and spatial transcriptomics technologies in gastric cancer (Review).Oncol Lett. 2024 Feb 14;27(4):152. doi: 10.3892/ol.2024.14285. eCollection 2024 Apr. Oncol Lett. 2024. PMID: 38406595 Free PMC article. Review.
-
Flexible parsing, interpretation, and editing of technical sequences with splitcode.Bioinformatics. 2024 Jun 3;40(6):btae331. doi: 10.1093/bioinformatics/btae331. Bioinformatics. 2024. PMID: 38876979 Free PMC article.
-
Flexible parsing, interpretation, and editing of technical sequences with splitcode.bioRxiv [Preprint]. 2023 Dec 9:2023.03.20.533521. doi: 10.1101/2023.03.20.533521. bioRxiv. 2023. Update in: Bioinformatics. 2024 Jun 3;40(6):btae331. doi: 10.1093/bioinformatics/btae331. PMID: 36993532 Free PMC article. Updated. Preprint.
References
-
- E. A. Winzeler, D. D. Shoemaker, A. Astromoff, H. Liang, K. Anderson, B. Andre, R. Bangham, R. Benito, J. D. Boeke, H. Bussey, A. M. Chu, C. Connelly, K. Davis, F. Dietrich, S. W. Dow, M. El Bakkoury, F. Foury, S. H. Friend, E. Gentalen, G. Giaever, J. H. Hegemann, T. Jones, M. Laub, H. Liao, N. Liebundguth, D. J. Lockhart, A. Lucau-Danila, M. Lussier, N. M’Rabet, P. Menard, M. Mittmann, C. Pai, C. Rebischung, J. L. Revuelta, L. Riles, C. J. Roberts, P. Ross-MacDonald, B. Scherens, M. Snyder, S. Sookhai-Mahadeo, R. K. Storms, S. Véronneau, M. Voet, G. Volckaert, T. R. Ward, R. Wysocki, G. S. Yen, K. Yu, K. Zimmermann, P. Philippsen, M. Johnston, R. W. Davis, Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999). - PubMed
-
- T. Roemer, J. Davies, G. Giaever, C. Nislow, Bugs, drugs and chemical genomics. Nat. Chem. Biol. 8, 46–56 (2011). - PubMed
-
- K. Berns, E. M. Hijmans, J. Mullenders, T. R. Brummelkamp, A. Velds, M. Heimerikx, R. M. Kerkhoven, M. Madiredjo, W. Nijkamp, B. Weigelt, R. Agami, W. Ge, G. Cavet, P. S. Linsley, R. L. Beijersbergen, R. Bernards, A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004). - PubMed
LinkOut - more resources
Full Text Sources

