Antiretroviral APOBEC3 cytidine deaminases alter HIV-1 provirus integration site profiles
- PMID: 36627271
- PMCID: PMC9832166
- DOI: 10.1038/s41467-022-35379-y
Antiretroviral APOBEC3 cytidine deaminases alter HIV-1 provirus integration site profiles
Abstract
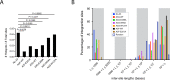
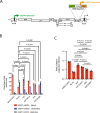
APOBEC3 (A3) proteins are host-encoded deoxycytidine deaminases that provide an innate immune barrier to retroviral infection, notably against HIV-1. Low levels of deamination are believed to contribute to the genetic evolution of HIV-1, while intense catalytic activity of these proteins can induce catastrophic hypermutation in proviral DNA leading to near-total HIV-1 restriction. So far, little is known about how A3 cytosine deaminases might impact HIV-1 proviral DNA integration sites in human chromosomal DNA. Using a deep sequencing approach, we analyze the influence of catalytic active and inactive APOBEC3F and APOBEC3G on HIV-1 integration site selections. Here we show that DNA editing is detected at the extremities of the long terminal repeat regions of the virus. Both catalytic active and non-catalytic A3 mutants decrease insertions into gene coding sequences and increase integration sites into SINE elements, oncogenes and transcription-silencing non-B DNA features. Our data implicates A3 as a host factor influencing HIV-1 integration site selection and also promotes what appears to be a more latent expression profile.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

