Engineering Cyborg Bacteria Through Intracellular Hydrogelation
- PMID: 36628538
- PMCID: PMC10037956
- DOI: 10.1002/advs.202204175
Engineering Cyborg Bacteria Through Intracellular Hydrogelation
Abstract
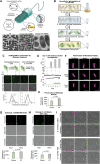
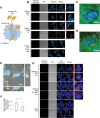
Natural and artificial cells are two common chassis in synthetic biology. Natural cells can perform complex tasks through synthetic genetic constructs, but their autonomous replication often causes safety concerns for biomedical applications. In contrast, artificial cells based on nonreplicating materials, albeit possessing reduced biochemical complexity, provide more defined and controllable functions. Here, for the first time, the authors create hybrid material-cell entities termed Cyborg Cells. To create Cyborg Cells, a synthetic polymer network is assembled inside each bacterium, rendering them incapable of dividing. Cyborg Cells preserve essential functions, including cellular metabolism, motility, protein synthesis, and compatibility with genetic circuits. Cyborg Cells also acquire new abilities to resist stressors that otherwise kill natural cells. Finally, the authors demonstrate the therapeutic potential by showing invasion into cancer cells. This work establishes a new paradigm in cellular bioengineering by exploiting a combination of intracellular man-made polymers and their interaction with the protein networks of living cells.
Keywords: cellular chassis; hybrid material; hydrogel; nonculturable cells; nonreplicating bacteria; synthetic biology.
© 2023 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
L.E.C‐L., C.M.H., and C.T submitted a provisional patent application covering the process described in this study. The rest of the authors declare no competing interests.
Figures



Similar articles
-
Fabrication of cyborg bacterial cells as living cell-material hybrids using intracellular hydrogelation.Nat Protoc. 2024 Dec;19(12):3613-3639. doi: 10.1038/s41596-024-01035-6. Epub 2024 Aug 22. Nat Protoc. 2024. PMID: 39174659 Free PMC article. Review.
-
Synthetic control of living cells by intracellular polymerization.Trends Biotechnol. 2024 Feb;42(2):241-252. doi: 10.1016/j.tibtech.2023.08.006. Epub 2023 Sep 23. Trends Biotechnol. 2024. PMID: 37743158 Free PMC article. Review.
-
Engineering Cyborg Pathogens through Intracellular Hydrogelation.ACS Synth Biol. 2024 Nov 15;13(11):3609-3620. doi: 10.1021/acssynbio.4c00420. Epub 2024 Oct 16. ACS Synth Biol. 2024. PMID: 39413025
-
Synthetic Biology-The Synthesis of Biology.Angew Chem Int Ed Engl. 2017 Jun 1;56(23):6396-6419. doi: 10.1002/anie.201609229. Epub 2017 Apr 25. Angew Chem Int Ed Engl. 2017. PMID: 27943572 Review.
-
A self-assembled dynamic extracellular matrix-like hydrogel system with multi-scale structures for cell bioengineering applications.Acta Biomater. 2023 May;162:211-225. doi: 10.1016/j.actbio.2023.03.015. Epub 2023 Mar 16. Acta Biomater. 2023. PMID: 36931420
Cited by
-
Fabrication of cyborg bacterial cells as living cell-material hybrids using intracellular hydrogelation.Nat Protoc. 2024 Dec;19(12):3613-3639. doi: 10.1038/s41596-024-01035-6. Epub 2024 Aug 22. Nat Protoc. 2024. PMID: 39174659 Free PMC article. Review.
-
Macrophage-Based Microrobots for Anticancer Therapy: Recent Progress and Future Perspectives.Biomimetics (Basel). 2023 Nov 18;8(7):553. doi: 10.3390/biomimetics8070553. Biomimetics (Basel). 2023. PMID: 37999194 Free PMC article. Review.
-
Synthetic control of living cells by intracellular polymerization.Trends Biotechnol. 2024 Feb;42(2):241-252. doi: 10.1016/j.tibtech.2023.08.006. Epub 2023 Sep 23. Trends Biotechnol. 2024. PMID: 37743158 Free PMC article. Review.
-
Combinatorial leaky probiotic for anticancer immunopotentiation and tumor eradication.Cell Rep Med. 2024 Nov 19;5(11):101793. doi: 10.1016/j.xcrm.2024.101793. Epub 2024 Oct 22. Cell Rep Med. 2024. PMID: 39442515 Free PMC article.
-
Environment signal dependent biocontainment systems for engineered organisms: Leveraging triggered responses and combinatorial systems.Synth Syst Biotechnol. 2024 Dec 20;10(2):356-364. doi: 10.1016/j.synbio.2024.12.005. eCollection 2025 Jun. Synth Syst Biotechnol. 2024. PMID: 39830078 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
