Comparison of transcriptome profiles between medulloblastoma primary and recurrent tumors uncovers novel variance effects in relapses
- PMID: 36635768
- PMCID: PMC9837941
- DOI: 10.1186/s40478-023-01504-1
Comparison of transcriptome profiles between medulloblastoma primary and recurrent tumors uncovers novel variance effects in relapses
Abstract
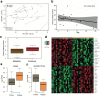
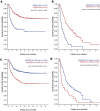
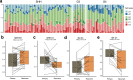
Nowadays medulloblastoma (MB) tumors can be treated with risk-stratified approaches with up to 80% success rate. However, disease relapses occur in approximately 30% of patients and successful salvage treatment strategies at relapse remain scarce. Acquired copy number changes or TP53 mutations are known to occur frequently in relapses, while methylation profiles usually remain highly similar to those of the matching primary tumors, indicating that in general molecular subgrouping does not change during the course of the disease. In the current study, we have used RNA sequencing data to analyze the transcriptome profiles of 43 primary-relapse MB pairs in order to identify specific molecular features of relapses within various tumor groups. Gene variance analysis between primary and relapse samples demonstrated the impact of age in SHH-MB: the changes in gene expression relapse profiles were more pronounced in the younger patients (< 10 years old), which were also associated with increased DNA aberrations and somatic mutations at relapse probably driving this effect. For Group 3/4 MB transcriptome data analysis uncovered clear sets of genes either active or decreased at relapse that are significantly associated with survival, thus could be potential predictive markers. In addition, deconvolution analysis of bulk transcriptome data identified progression-associated differences in cell type enrichment. The proportion of undifferentiated progenitors increased in SHH-MB relapses with a concomitant decrease of differentiated neuron-like cells, while in Group 3/4 MB relapses cell cycle activity increases and differentiated neuron-like cells proportion decreases as well. Thus, our findings uncovered significant transcriptome changes in the molecular signatures of relapsed MB and could be potentially useful for further clinical purposes.
Keywords: Medulloblastoma; Prognosis; Relapses; Transcriptomics.
© 2023. The Author(s).
Conflict of interest statement
The authors have no competing interests.
Figures


References
-
- Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman W-H, Pagès F, Trajanoski Z, Galon J. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics. 2009;25:1091–1093. doi: 10.1093/bioinformatics/btp101. - DOI - PMC - PubMed
-
- Dobin A, Gingeras TR. Data mining techniques for the life sciences. Springer; 2016. Optimizing RNA-Seq mapping with STAR; pp. 245–262. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

