AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots
- PMID: 36644757
- PMCID: PMC9832943
- DOI: 10.1093/synbio/ysac032
AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots
Abstract
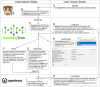
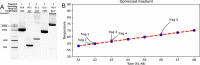
As one of the newest fields of engineering, synthetic biology relies upon a trial-and-error Design-Build-Test-Learn (DBTL) approach to simultaneously learn how a function is encoded in biology and attempt to engineer it. Many software and hardware platforms have been developed to automate, optimize and algorithmically perform each step of the DBTL cycle. However, there are many fewer options for automating the build step. Build typically involves deoxyribonucleic acid (DNA) assembly, which remains manual, low throughput and unreliable in most cases and limits our ability to advance the science and engineering of biology. Here, we present AssemblyTron, an open-source Python package to integrate j5 DNA assembly design software outputs with build implementation in Opentrons liquid handling robotics with minimal human intervention. We demonstrate the versatility of AssemblyTron through several scarless, multipart DNA assemblies, beginning from fragment amplification. We show that AssemblyTron can perform polymerase chain reactions across a range of fragment lengths and annealing temperatures by using an optimal annealing temperature gradient calculation algorithm. We then demonstrate that AssemblyTron can perform Golden Gate and homology-dependent in vivo assemblies (IVAs) with comparable fidelity to manual assemblies by simultaneously building four four-fragment assemblies of chromoprotein reporter expression plasmids. Finally, we used AssemblyTron to perform site-directed mutagenesis reactions via homology-dependent IVA also achieving comparable fidelity to manual assemblies as assessed by sequencing. AssemblyTron can reduce the time, training, costs and wastes associated with synthetic biology, which, along with open-source and affordable automation, will further foster the accessibility of synthetic biology and accelerate biological research and engineering.
Keywords: DNA assembly; design cycle; laboratory automation; molecular cloning; robotic liquid handling.
© The Author(s) 2023. Published by Oxford University Press.
Figures




Similar articles
-
Biofoundry-Assisted Golden Gate Cloning with AssemblyTron.Methods Mol Biol. 2025;2850:133-147. doi: 10.1007/978-1-0716-4220-7_8. Methods Mol Biol. 2025. PMID: 39363070
-
Standardizing Automated DNA Assembly: Best Practices, Metrics, and Protocols Using Robots.SLAS Technol. 2019 Jun;24(3):282-290. doi: 10.1177/2472630318825335. Epub 2019 Feb 15. SLAS Technol. 2019. PMID: 30768372 Free PMC article.
-
basicsynbio and the BASIC SEVA collection: software and vectors for an established DNA assembly method.Synth Biol (Oxf). 2022 Oct 11;7(1):ysac023. doi: 10.1093/synbio/ysac023. eCollection 2022. Synth Biol (Oxf). 2022. PMID: 36381610 Free PMC article.
-
Accelerating strain engineering in biofuel research via build and test automation of synthetic biology.Curr Opin Biotechnol. 2021 Feb;67:88-98. doi: 10.1016/j.copbio.2021.01.010. Epub 2021 Jan 25. Curr Opin Biotechnol. 2021. PMID: 33508635 Review.
-
Physical Laboratory Automation in Synthetic Biology.ACS Synth Biol. 2023 Nov 17;12(11):3156-3169. doi: 10.1021/acssynbio.3c00345. Epub 2023 Nov 7. ACS Synth Biol. 2023. PMID: 37935025 Review.
Cited by
-
Directed assembly of single-stranded DNA fragments for data storage via protein-free catalytic splint ligation.Nucleic Acids Res. 2025 Jun 20;53(12):gkaf582. doi: 10.1093/nar/gkaf582. Nucleic Acids Res. 2025. PMID: 40586308 Free PMC article.
-
TidyTron: Reducing lab waste using validated wash-and-reuse protocols for common plasticware in Opentrons OT-2 lab robots.SLAS Technol. 2024 Apr;29(2):100107. doi: 10.1016/j.slast.2023.08.007. Epub 2023 Sep 9. SLAS Technol. 2024. PMID: 37696493 Free PMC article.
-
Getting the Right Clones in an Automated Manner: An Alternative to Sophisticated Colony-Picking Robotics.Bioengineering (Basel). 2024 Sep 3;11(9):892. doi: 10.3390/bioengineering11090892. Bioengineering (Basel). 2024. PMID: 39329634 Free PMC article.
-
The knowledge driven DBTL cycle provides mechanistic insights while optimising dopamine production in Escherichia coli.Microb Cell Fact. 2025 May 16;24(1):111. doi: 10.1186/s12934-025-02729-6. Microb Cell Fact. 2025. PMID: 40380156 Free PMC article.
-
Semi-automated workflow for high-throughput Agrobacterium-mediated plant transformation.Plant J. 2025 Apr;122(1):e70118. doi: 10.1111/tpj.70118. Plant J. 2025. PMID: 40220013 Free PMC article.
References
-
- Casini A., Storch M., Baldwin G.S. and Ellis T. (2015) Bricks and blueprints: methods and standards for DNA assembly. Nat. Rev. Mol. Cell Biol., 16, 568–576. - PubMed
-
- Appleton E., Densmore D., Madsen C. and Roehner N. (2017) Needs and opportunities in bio-design automation: four areas for focus. Curr. Opin. Chem. Biol., 40, 111–118. - PubMed
-
- Li M.Z., Elledge S.J. (2012) SLIC: a method for sequence- and ligation-independent cloning. In: Peccoud J. (ed). Gene Synthesis. Methods in Molecular Biology, Vol. 852. Humana Press, Totowa, NJ, pp. 51–59. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
