Cell type characterization of spatiotemporal gene co-expression modules in Down syndrome brain
- PMID: 36647384
- PMCID: PMC9840153
- DOI: 10.1016/j.isci.2022.105884
Cell type characterization of spatiotemporal gene co-expression modules in Down syndrome brain
Abstract
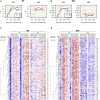
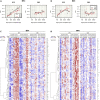
Down syndrome (DS) is the most common genetic cause of intellectual disability and increases the risk of other brain-related dysfunctions, like seizures, early-onset Alzheimer's disease, and autism. To reveal the molecular profiles of DS-associated brain phenotypes, we performed a meta-data analysis of the developmental DS brain transcriptome at cell type and co-expression module levels. In the DS brain, astrocyte-, microglia-, and endothelial cell-associated genes show upregulated patterns, whereas neuron- and oligodendrocyte-associated genes show downregulated patterns. Weighted gene co-expression network analysis identified cell type-enriched co-expressed gene modules. We present eight representative cell-type modules for neurons, astrocytes, oligodendrocytes, and microglia. We classified the neuron modules into glutamatergic and GABAergic neurons and associated them with detailed subtypes. Cell type modules were interpreted by analyzing spatiotemporal expression patterns, functional annotations, and co-expression networks of the modules. This study provides insight into the mechanisms underlying brain abnormalities in DS and related disorders.
Keywords: Developmental neuroscience; Transcriptomics.
© 2022 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures










Similar articles
-
Cell-Type-Specific Gene Modules Related to the Regional Homogeneity of Spontaneous Brain Activity and Their Associations With Common Brain Disorders.Front Neurosci. 2021 Apr 20;15:639527. doi: 10.3389/fnins.2021.639527. eCollection 2021. Front Neurosci. 2021. PMID: 33958982 Free PMC article.
-
Co-expression network analysis of Down's syndrome based on microarray data.Exp Ther Med. 2016 Sep;12(3):1503-1508. doi: 10.3892/etm.2016.3462. Epub 2016 Jun 17. Exp Ther Med. 2016. PMID: 27588071 Free PMC article.
-
SLC9A9 Co-expression modules in autism-associated brain regions.Autism Res. 2017 Mar;10(3):414-429. doi: 10.1002/aur.1670. Epub 2016 Jul 21. Autism Res. 2017. PMID: 27439572
-
Systematic Analysis of the Transcriptome Profiles and Co-Expression Networks of Tumour Endothelial Cells Identifies Several Tumour-Associated Modules and Potential Therapeutic Targets in Hepatocellular Carcinoma.Cancers (Basel). 2021 Apr 7;13(8):1768. doi: 10.3390/cancers13081768. Cancers (Basel). 2021. PMID: 33917186 Free PMC article.
-
Astrocytes in Down Syndrome Across the Lifespan.Front Cell Neurosci. 2021 Aug 18;15:702685. doi: 10.3389/fncel.2021.702685. eCollection 2021. Front Cell Neurosci. 2021. PMID: 34483840 Free PMC article. Review.
Cited by
-
A review of multi-omics data integration through deep learning approaches for disease diagnosis, prognosis, and treatment.Front Genet. 2023 Jul 20;14:1199087. doi: 10.3389/fgene.2023.1199087. eCollection 2023. Front Genet. 2023. PMID: 37547471 Free PMC article. Review.
-
Differential Roles of Astrocytic CSF1 in Alzheimer's Disease and Cerebral Amyloid Angiopathy: Insights from Transcriptomic Analysis.Aging Dis. 2024 Nov 5;16(5):3137-3153. doi: 10.14336/AD.2024.10530. Aging Dis. 2024. PMID: 39571157 Free PMC article.
-
Brain Mitochondrial Bioenergetics in Genetic Neurodevelopmental Disorders: Focus on Down, Rett and Fragile X Syndromes.Int J Mol Sci. 2023 Aug 6;24(15):12488. doi: 10.3390/ijms241512488. Int J Mol Sci. 2023. PMID: 37569863 Free PMC article. Review.
-
Early life phenobarbital exposure dysregulates the hippocampal transcriptome.Front Pharmacol. 2024 Mar 28;15:1340691. doi: 10.3389/fphar.2024.1340691. eCollection 2024. Front Pharmacol. 2024. PMID: 38606173 Free PMC article.
-
Beyond Quiescent and Active: Intermediate Microglial Transcriptomic States in a Mouse Model of Down Syndrome.Int J Mol Sci. 2024 Mar 14;25(6):3289. doi: 10.3390/ijms25063289. Int J Mol Sci. 2024. PMID: 38542263 Free PMC article.
References
LinkOut - more resources
Full Text Sources

