Small RNA-mediated DNA methylation during plant reproduction
- PMID: 36651080
- PMCID: PMC10226566
- DOI: 10.1093/plcell/koad010
Small RNA-mediated DNA methylation during plant reproduction
Abstract
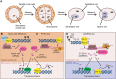
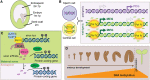
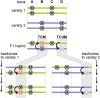
Reproductive tissues are a rich source of small RNAs, including several classes of short interfering (si)RNAs that are restricted to this stage of development. In addition to RNA polymerase IV-dependent 24-nt siRNAs that trigger canonical RNA-directed DNA methylation, abundant reproductive-specific siRNAs are produced from companion cells adjacent to the developing germ line or zygote and may move intercellularly before inducing methylation. In some cases, these siRNAs are produced via non-canonical biosynthesis mechanisms or from sequences with little similarity to transposons. While the precise role of these siRNAs and the methylation they trigger is unclear, they have been implicated in specifying a single megaspore mother cell, silencing transposons in the male germ line, mediating parental dosage conflict to ensure proper endosperm development, hypermethylation of mature embryos, and trans-chromosomal methylation in hybrids. In this review, we summarize the current knowledge of reproductive siRNAs, including their biosynthesis, transport, and function.
© American Society of Plant Biologists 2023. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Conflict of interest statement
Conflict of interest statement. None declared.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

