PhaTYP: predicting the lifestyle for bacteriophages using BERT
- PMID: 36659812
- PMCID: PMC9851330
- DOI: 10.1093/bib/bbac487
PhaTYP: predicting the lifestyle for bacteriophages using BERT
Abstract
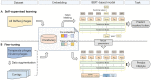
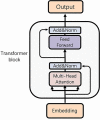
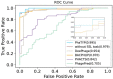
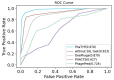
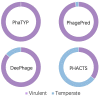
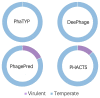
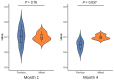
Bacteriophages (or phages), which infect bacteria, have two distinct lifestyles: virulent and temperate. Predicting the lifestyle of phages helps decipher their interactions with their bacterial hosts, aiding phages' applications in fields such as phage therapy. Because experimental methods for annotating the lifestyle of phages cannot keep pace with the fast accumulation of sequenced phages, computational method for predicting phages' lifestyles has become an attractive alternative. Despite some promising results, computational lifestyle prediction remains difficult because of the limited known annotations and the sheer amount of sequenced phage contigs assembled from metagenomic data. In particular, most of the existing tools cannot precisely predict phages' lifestyles for short contigs. In this work, we develop PhaTYP (Phage TYPe prediction tool) to improve the accuracy of lifestyle prediction on short contigs. We design two different training tasks, self-supervised and fine-tuning tasks, to overcome lifestyle prediction difficulties. We rigorously tested and compared PhaTYP with four state-of-the-art methods: DeePhage, PHACTS, PhagePred and BACPHLIP. The experimental results show that PhaTYP outperforms all these methods and achieves more stable performance on short contigs. In addition, we demonstrated the utility of PhaTYP for analyzing the phage lifestyle on human neonates' gut data. This application shows that PhaTYP is a useful means for studying phages in metagenomic data and helps extend our understanding of microbial communities.
Keywords: BRET; deep learning; phage lifestyle prediction; virulent and temperate phages.
© The Author(s) 2022. Published by Oxford University Press.
Figures





References
-
- McGrath S, van Sinderen, et al. Bacteriophage: Genetics and Molecular Biology. Wymondham, UK: Caister Academic Press, 2007.
-
- Moineau S. Applications of phage resistance in lactic acid bacteria. Lactic Acid Bact 1999;76(1–4):377–82. - PubMed
-
- Brüssow H, Desiere F. Comparative phage genomics and the evolution of siphoviridae: insights from dairy phages. Mol Microbiol 2001;39(2):213–23. - PubMed

